- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- For Contributors
- Policies
- E-Submission

- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- Policies
- For Contributors
Articles
- Page Path
- HOME > J. Microbiol > Volume 64(5); 2026 > Article
-
Full article
Crystal structure of Bcl-2 from lymphocystis disease virus 2 in complex with the BH3 domain of zebrafish BaxA -
Dahwan Lim1,2,†, So Hyeon Park1,3,†, Joon Sig Choi3, Ho-Chul Shin4, Seung Jun Kim1,4,*, Bonsu Ku1,*

-
Journal of Microbiology 2026;64(5):e2512006.
DOI: https://doi.org/10.71150/jm.2512006
Published online: April 23, 2026
1Orphan Disease Therapeutic Target Research Center, Korea Research Institute of Bioscience and Biotechnology, Daejeon 34141, Republic of Korea
2Infectious Diseases Therapeutic Research Center, Korea Research Institute of Chemical Technology, Daejeon 34114, Republic of Korea
3Department of Biochemistry, Chungnam National University, Daejeon 34134, Republic of Korea
4Critical Diseases Diagnostics Convergence Research Center, Korea Research Institute of Bioscience and Biotechnology, Daejeon 34141, Republic of Korea
- *Correspondence Seung Jun Kim ksj@kribb.re.kr Bonsu Ku bku@kribb.re.kr
- †These authors contributed equally to this work.
© The Microbiological Society of Korea
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
- 733 Views
- 22 Download
ABSTRACT
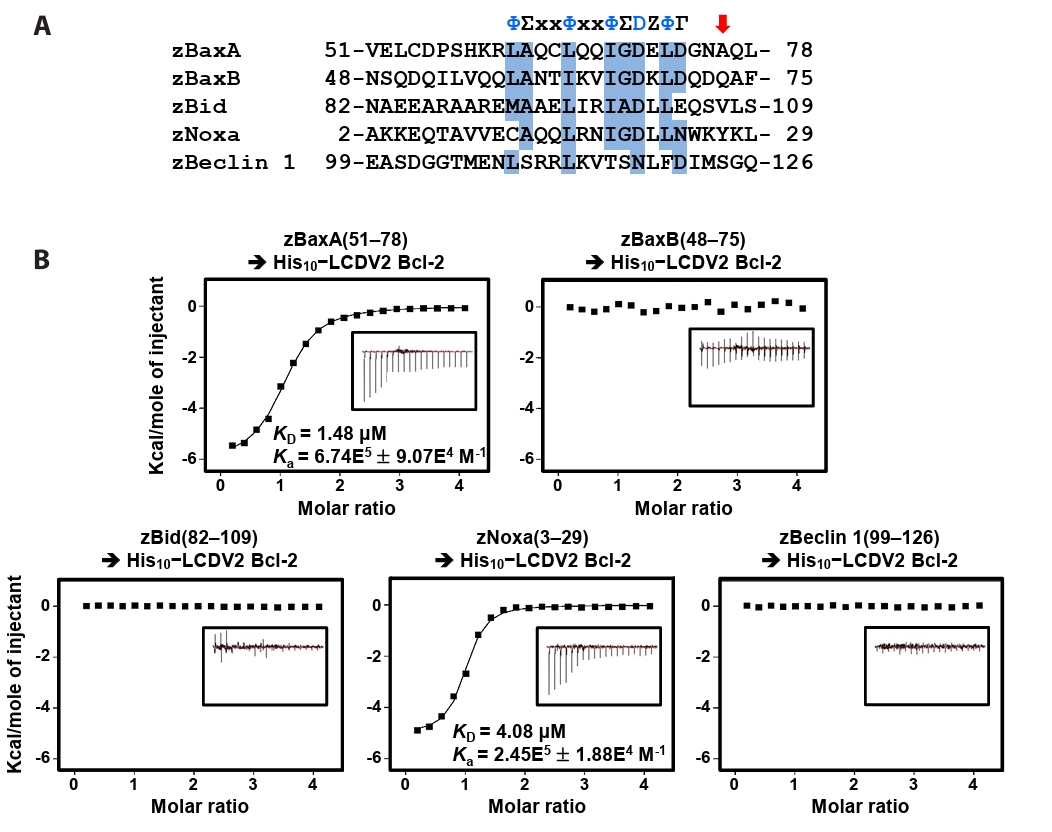
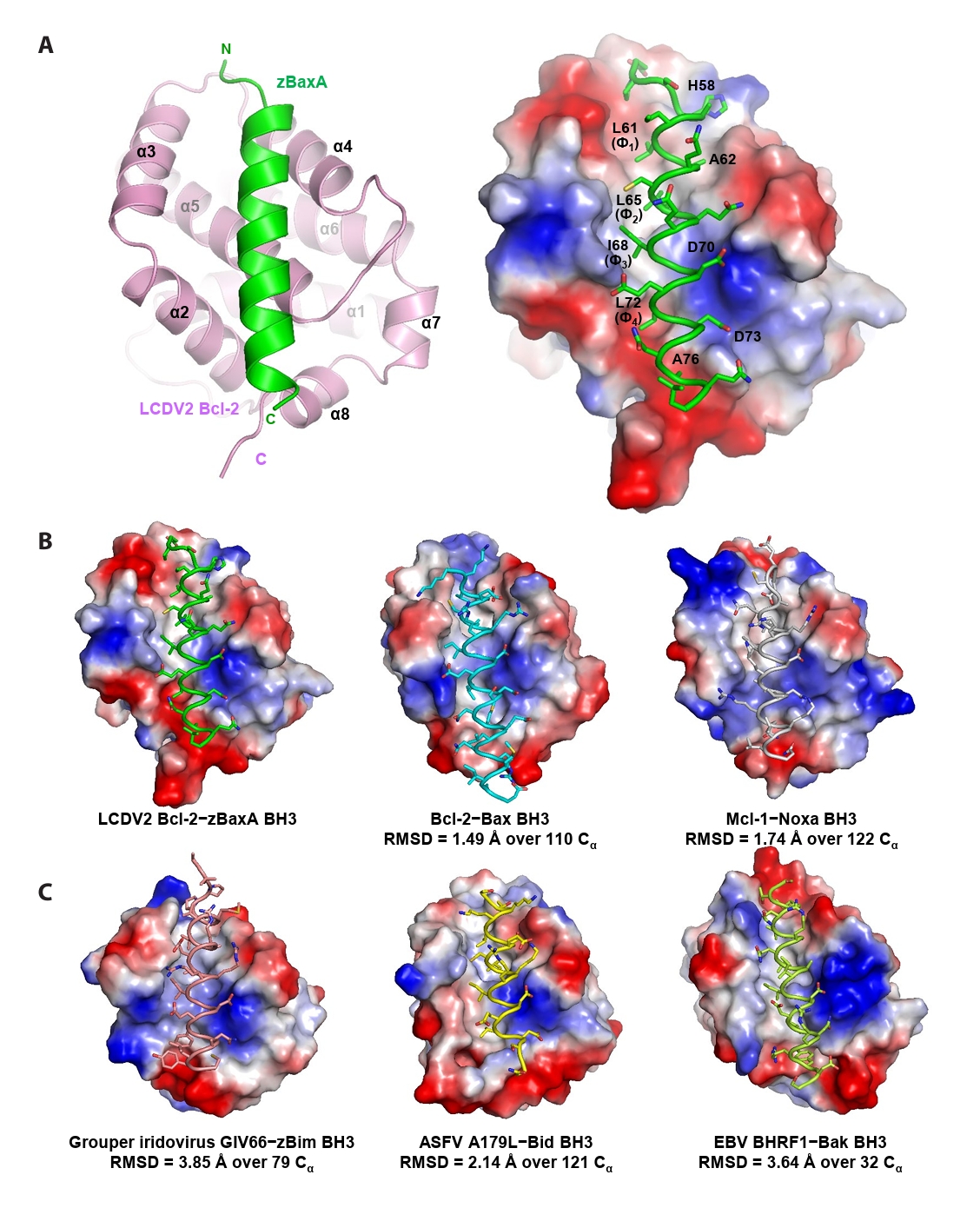
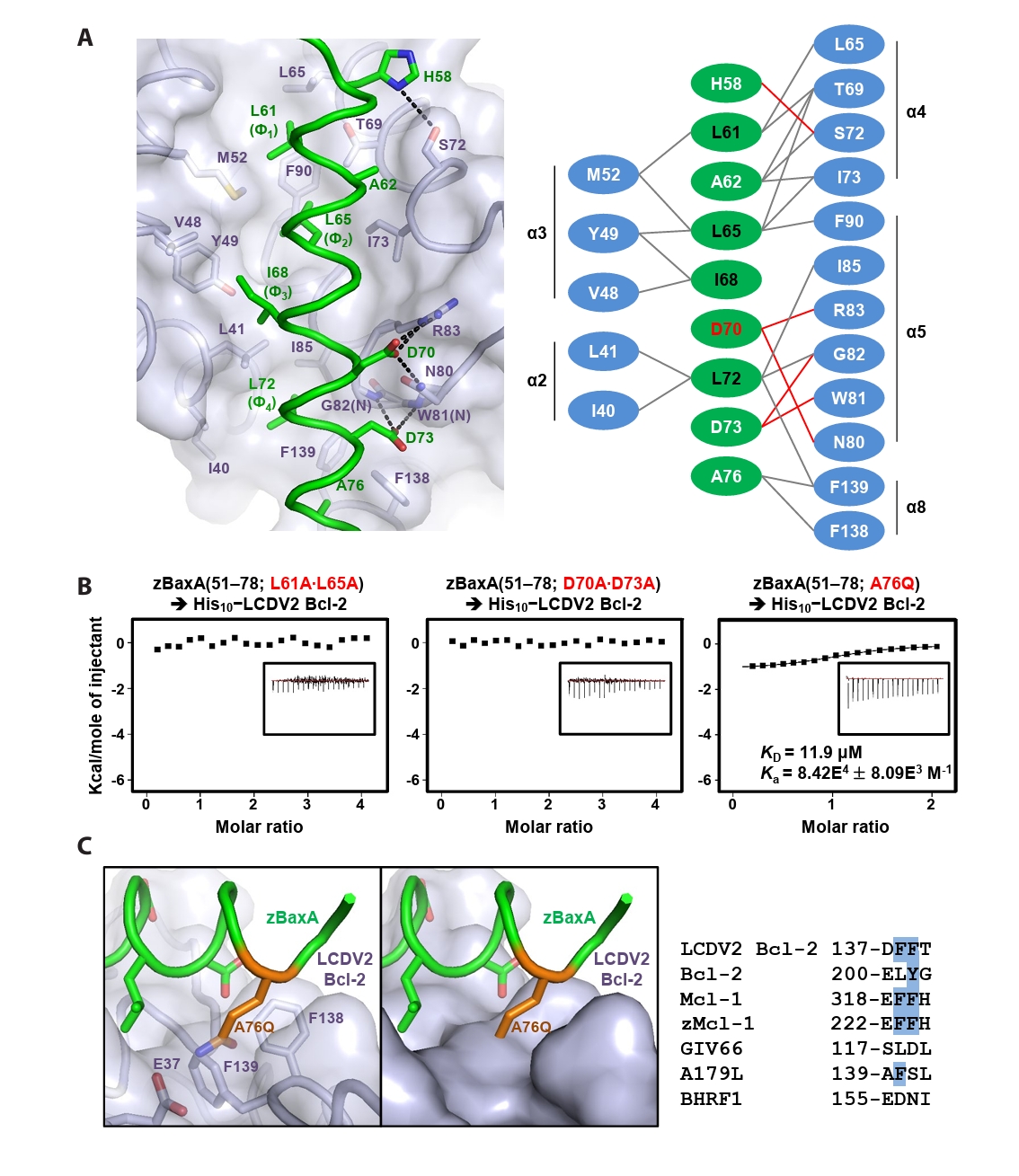
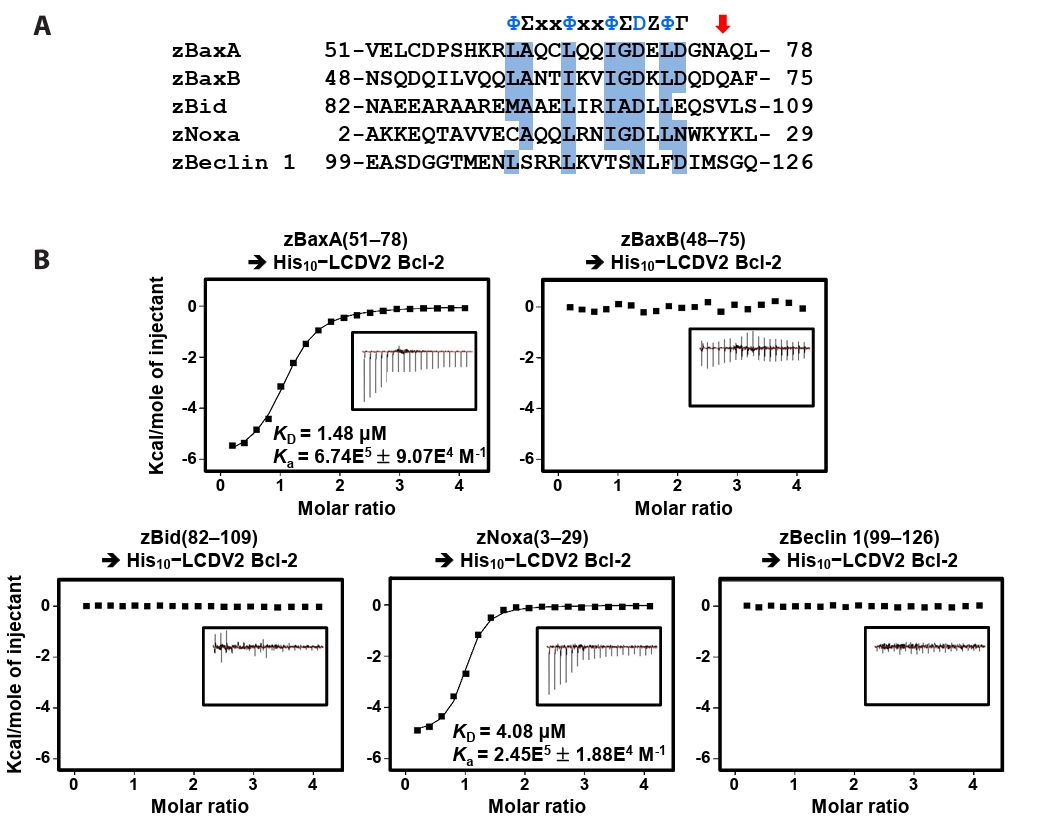
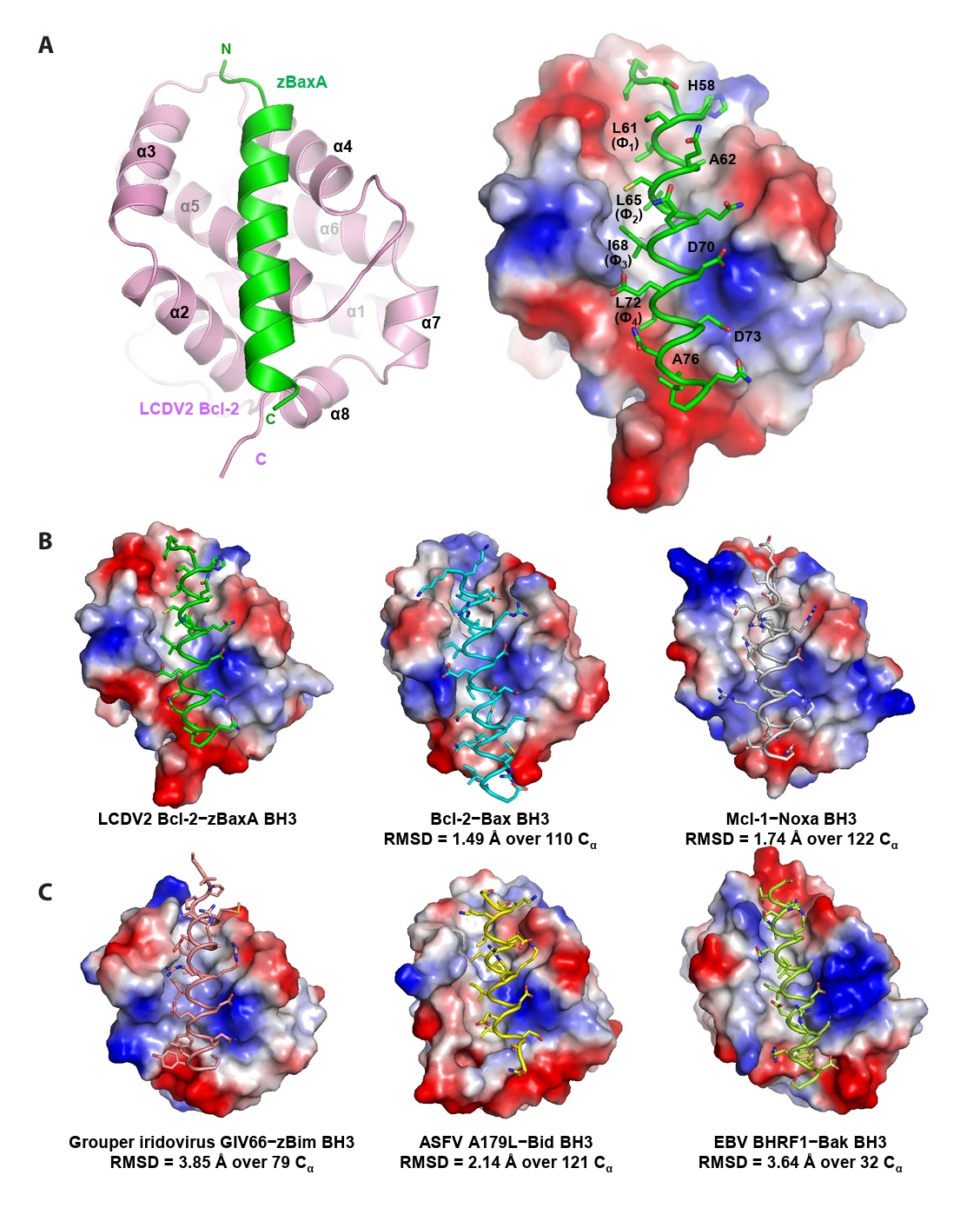
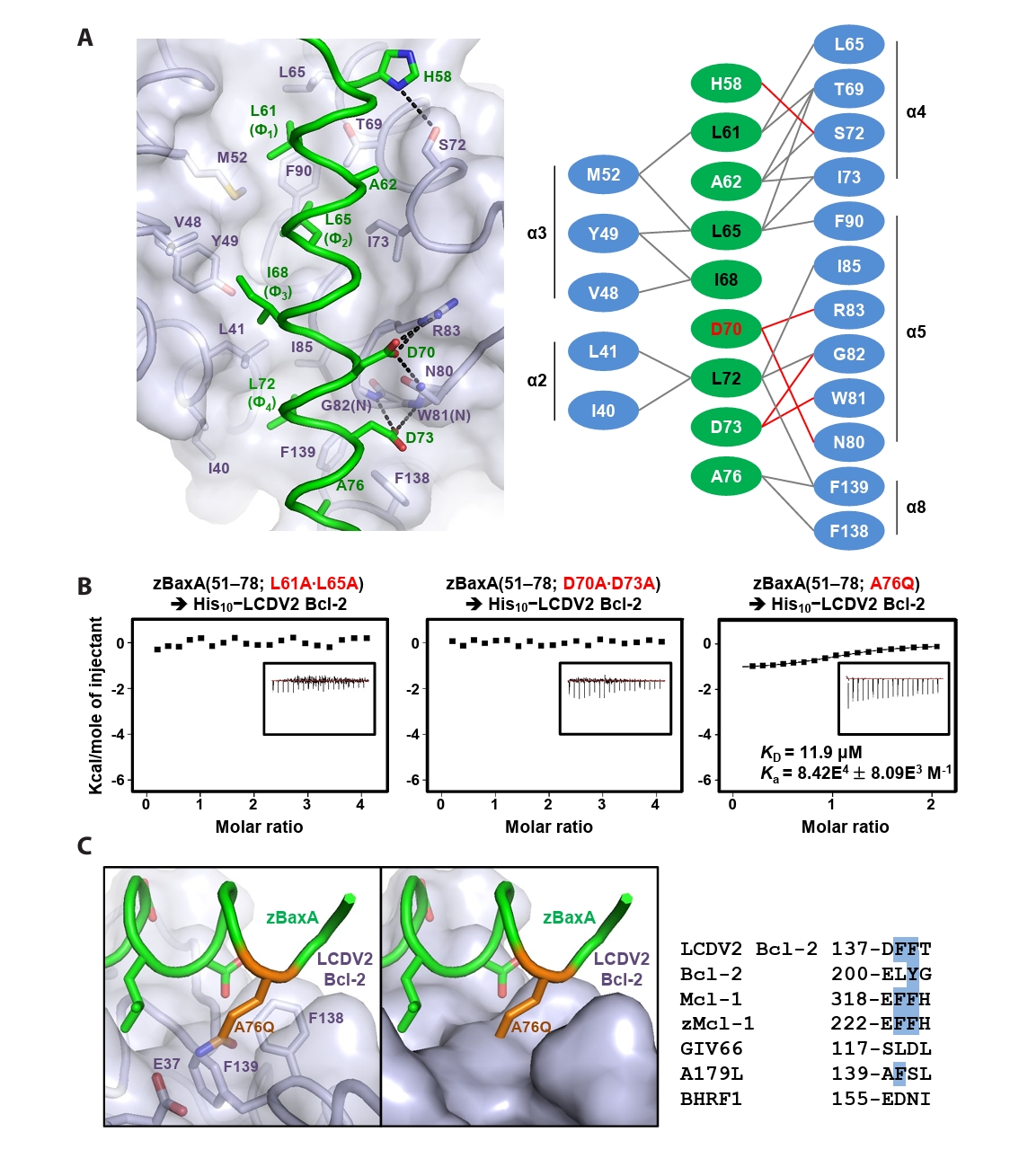
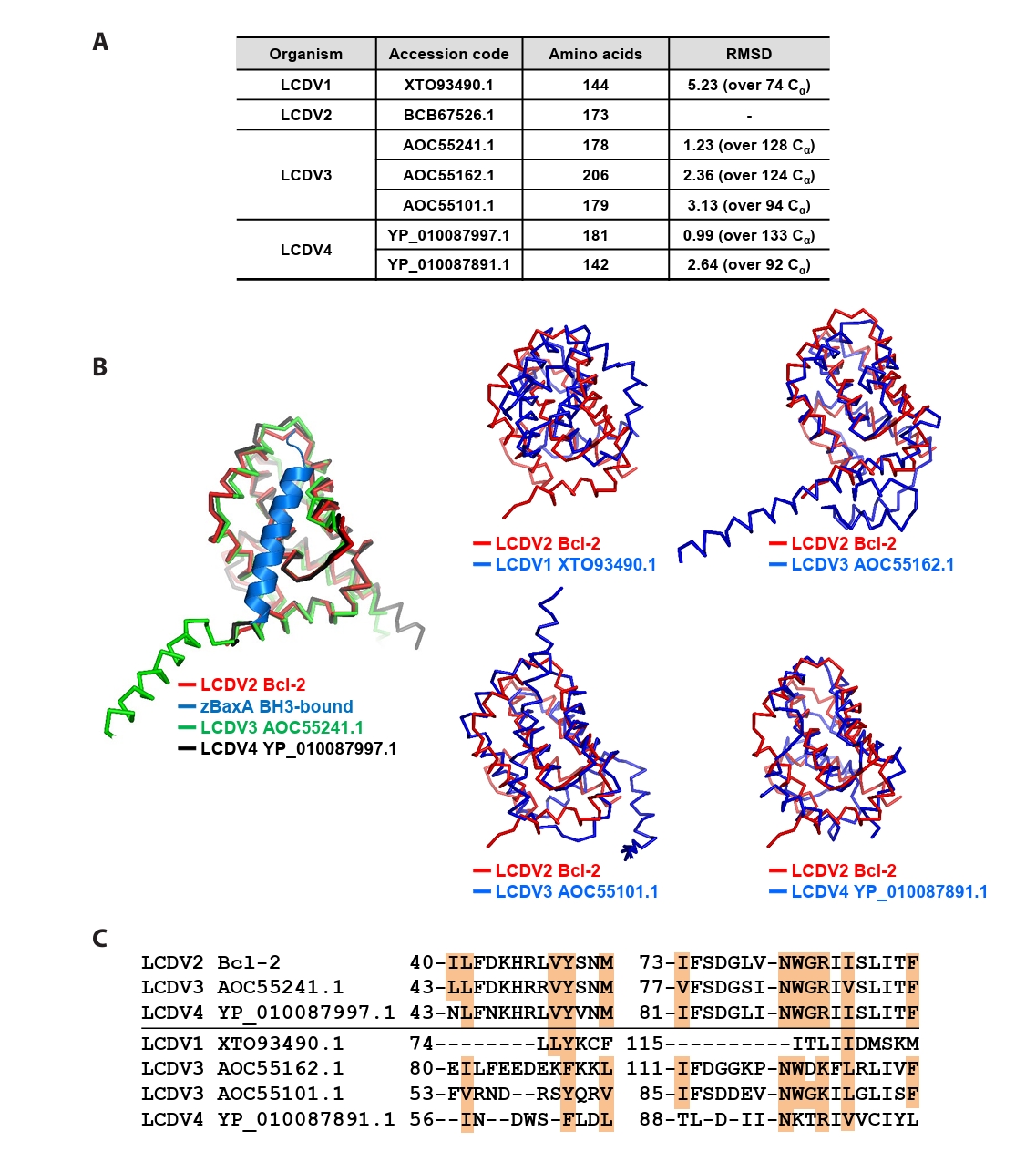
- Lymphocystis disease viruses (LCDVs), members of the Lymphocystivirus genus of the Iridoviridae family, infect various freshwater and marine fish species. They cause the chronic disease lymphocystis, which is non-fatal, but substantially reduces the commercial value of the infected fish. To date, four genotypes of LCDV (LCDV1–4) have been identified, all of which encode the viral homologue of B-cell lymphoma 2 (Bcl-2), a key inhibitor of apoptosis. In this study, we performed biochemical and structural analyses of LCDV2 Bcl-2. Binding assays revealed that LCDV2 Bcl-2 exhibits binding selectivity toward BH3 domain-containing zebrafish proteins. It interacted with zBaxA and zNoxa, but not with zBaxB, zBid, or zBeclin 1, distinguishing it from mammalian and herpesviral Bcl-2 proteins. Subsequent structural determination of LCDV2 Bcl-2 in complex with the BH3 domain of zBaxA demonstrated that they interact in a canonical manner, primarily mediated by the BH3 consensus motif residues of zBaxA. In addition, a subpocket formed by two phenylalanine residues in LCDV2 Bcl-2 plays a key role in determining binding selectivity.
Introduction
Materials and Methods
Results and Discussion
Conclusion
Acknowledgments
We are grateful to Dr. Eunha Hwang (Korea Basic Science Institute, Korea) for the assistance with ITC experiments. This study used Beamline 11C at the Pohang Accelerator Laboratory in Korea. This work was supported by grants from the National Research Foundation of Korea (RS-2023-00278696 and RS-2024-00348679), National Research Council of Science and Technology (CRC22021-700), and Korea Research Institute of Bioscience and Biotechnology Research Initiative Programs (KGM9952623 and KQM0052611), supervised by the Ministry of Science and ICT, Korea.
Conflict of Interest
The authors declare that they have no competing interests.
Data Availability
The coordinates of the LCDV2 Bcl-2−zBaxA(51–78) structure, together with the structure factors, were deposited in the Protein Data Bank under accession code 21CP.
| Protein | LCDV2 Bcl-2−zBaxA(51–78) |
|---|---|
| Data collection | |
| Space group | P41212 |
| Unit cell dimensions | |
| a, b, c (Å) / a = β = γ (°) | 89.14, 89.14, 50.60 / 90 |
| Resolution (Å) | 50.0−2.3 (2.34−2.30)a |
| Rsymb (%) | 11.6 (30.1) |
| I / σ (I) | 26.3 (3.9) |
| Completeness (%) | 96.8 (94.6) |
| Redundancy | 5.5 |
| Refinement | |
| Resolution (Å) | 50.0−2.3 |
| Number of reflections | 9304 |
| Rworkc / Rfree (%) | 19.3 / 24.3 |
| Number of atoms | |
| Protein | 1133 |
| Peptide | 176 |
| Water | 28 |
| RMSD | |
| Bond lengths (Å) | 0.006 |
| Bond angles (°) | 0.728 |
| Ramachandran plot (%) | |
| Most favored region | 96.9 |
| Additionally allowed region | 3.1 |
| Average B-values (Å2) | |
| Protein | 43.5 |
| Peptide | 46.5 |
| Water | 45.6 |
aThe numbers in parentheses are statistics from the shell with the highest resolution.
bRsym = Σ |Iobs - Iavg| / Iobs, where Iobs is the observed intensity of individual reflection and Iavg is the average across symmetry equivalents.
cRwork = Σ ||Fo| - |Fc|| / Σ |Fo|, where |Fo| and |Fc| are the observed and calculated structure factor amplitudes, respectively. Rfree was calculated with 10.0% of the data.
- Adams PD, Afonine PV, Bunkoczi G, Chen VB, Davis IW, et al. 2010. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr D Biol Crystallogr. 66: 213–221. ArticlePubMedPMC
- Aguilar F, Yu S, Grant RA, Swanson S, Ghose D, et al. 2023. Peptides from human BNIP5 and PXT1 and non-native binders of pro-apoptotic BAK can directly activate or inhibit BAK-mediated membrane permeabilization. Structure. 31: 265–281.e7. ArticlePubMedPMC
- Banjara S, Caria S, Dixon LK, Hinds MG, Kvansakul M. 2017. Structural insight into African swine fever virus A179L-mediated inhibition of apoptosis. J Virol. 91: e02228–16. ArticlePubMedPMCLink
- Banjara S, Mao J, Ryan TM, Caria S, Kvansakul M. 2018. Grouper iridovirus GIV66 is a Bcl-2 protein that inhibits apoptosis by exclusively sequestering Bim. J Biol Chem. 293: 5464–5477. ArticlePubMedPMC
- Benkaroun J, Bergmann SM, Romer-Oberdorfer A, Demircan MD, Tamer C, et al. 2022. New insights into lymphocystis disease virus genome diversity. Viruses. 14: 2741.ArticlePubMedPMC
- Croce CM, Vaux D, Strasser A, Opferman JT, Czabotar PE, et al. 2025. The BCL-2 protein family: from discovery to drug development. Cell Death Differ. 32: 1369–1381. ArticlePubMedPMCPDF
- Czabotar PE, Lee EF, van Delft MF, Day CL, Smith BJ, et al. 2007. Structural insights into the degradation of Mcl-1 induced by BH3 domains. Proc Natl Acad Sci USA. 104: 6217–6222. ArticlePubMedPMC
- Doszpoly A, Kajan GL, Puentes R, Perretta A. 2020. Complete genome sequence and analysis of a novel lymphocystivirus detected in whitemouth croaker (Micropogonias furnieri): lymphocystis disease virus 4. Arch Virol. 165: 1215–1218. ArticlePubMedPMCPDF
- Emsley P, Cowtan K. 2004. Coot: model-building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr. 60: 2126–2132. ArticlePubMed
- Jung S, Lim D, Choi JS, Shin HC, Kim SJ, et al. 2025. Crystal structures of the μ2 subunit of clathrin-adaptor protein 2 in complex with peptides derived from human papillomavirus 16 E7. J Microbiol. 63: e2505003. ArticlePubMedPDF
- Kawato S, Nozaki R, Hirono I, Kondo H. 2021. Genome sequence of lymphocystis disease virus 2 LCDV-JP_Oita_2018, isolated from a diseased Japanese flounder (Paralichthys olivaceus) in Japan. Microbiol Resour Announc. 10: e00547-21. ArticlePubMedPMCLink
- Kratz E, Eimon PM, Mukhyala K, Stern H, Zha J, et al. 2006. Functional characterization of the Bcl-2 gene family in the zebrafish. Cell Death Differ. 13: 1631–1640. ArticlePubMedPDF
- Ku B. 2023. Structural analysis of the interaction between Bcl-xL and the noncanonical BH3 domain of non-Bcl-2 family proteins. Curr Protein Pept Sci. 24: 296–306. ArticlePubMedLink
- Ku B, Liang C, Jung JU, Oh BH. 2011. Evidence that inhibition of BAX activation by BCL-2 involves its tight and preferential interaction with the BH3 domain of BAX. Cell Res. 21: 627–641. ArticlePubMedPDF
- Ku B, Woo JS, Liang C, Lee KH, Hong HS, et al. 2008. Structural and biochemical bases for the inhibition of autophagy and apoptosis by viral BCL-2 of murine γ-herpesvirus 68. PLoS Pathog. 4: e25. ArticlePubMedPMC
- Kvansakul M, Hinds MG. 2013. Structural biology of the Bcl-2 family and its mimicry by viral proteins. Cell Death Dis. 4: e909. ArticlePubMedPMCPDF
- Kvansakul M, Hinds MG. 2015. The Bcl-2 family: structures, interactions and targets for drug discovery. Apoptosis. 20: 136–150. ArticlePubMedPDF
- Kvansakul M, Wei AH, Fletcher JI, Willis SN, Chen L, et al. 2010. Structural basis for apoptosis inhibition by Epstein-Barr virus BHRF1. PLoS Pathog. 6: e1001236. ArticlePubMedPMC
- Kvansakul M, Yang H, Fairlie WD, Czabotar PE, Fischer SF, et al. 2008. Vaccinia virus anti-apoptotic F1L is a novel Bcl-2-like domain-swapped dimer that binds a highly selective subset of BH3-containing death ligands. Cell Death Differ. 15: 1564–1571. ArticlePubMedPDF
- Leiva-Rebollo R, Labella AM, Gemez-Mata J, Castro D, Borrego JJ. 2024. Fish Iridoviridae : infection, vaccination and immune response. Vet Res. 55: 88.ArticlePubMedPMCPDF
- Levine B, Sinha S, Kroemer G. 2008. Bcl-2 family members: dual regulators of apoptosis and autophagy. Autophagy. 4: 600–606. ArticlePubMedPMC
- Lim D, Choe SH, Jin S, Lee S, Kim Y, et al. 2023. Structural basis for proapoptotic activation of Bak by the noncanonical BH3-only protein Pxt1. PLoS Biol. 21: e3002156. ArticlePubMedPMC
- Lim D, Jeong DE, Shin HC, Choi JS, Seo J, et al. 2024. Crystal structure of Bak bound to the BH3 domain of Bnip5, a noncanonical BH3 domain-containing protein. Proteins. 92: 44–51. ArticlePubMed
- Lim D, Shin HC, Choi JS, Kim SJ, Ku B. 2021. Crystal structure of human LC8 bound to a peptide from Ebola virus VP35. J Microbiol. 59: 410–416. ArticlePubMedPDF
- Lin PW, Huang YJ, John JA, Chang YN, Yuan CH, et al. 2008. Iridovirus Bcl-2 protein inhibits apoptosis in the early stage of viral infection. Apoptosis. 13: 165–176. ArticlePubMedPDF
- López-Bueno A, Mavian C, Labella AM, Castro D, Borrego JJ, et al. 2016. Concurrence of iridovirus, polyomavirus, and a unique member of a new group of fish papillomaviruses in lymphocystis disease-affected gilthead sea bream. J Virol. 90: 8768–8779. ArticlePubMedPMCLink
- McCoy AJ, Grosse-Kunstleve RW, Adams PD, Winn MD, Storoni LC, et al. 2007. Phaser crystallographic software. J Appl Crystallogr. 40: 658–674. ArticlePubMedPMC
- Otwinowski Z, Minor W. 1997. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276: 307–326. PubMed
- Sasaki T, Lian S, Qi J, Bayliss PE, Carr CE, et al. 2014. Aberrant autolysosomal regulation is linked to the induction of embryonic senescence: differential roles of Beclin 1 and p53 in vertebrate Spns1 deficiency. PLoS Genet. 10: e1004409. ArticlePubMedPMC
- Singh R, Letai A, Sarosiek K. 2019. Regulation of apoptosis in health and disease: the balancing act of BCL-2 family proteins. Nat Rev Mol Cell Biol. 20: 175–193. ArticlePubMedPMCPDF
- Suraweera CD, Espinoza B, Hinds MG, Kvansakul M. 2024. Mastering death: the roles of viral Bcl-2 in dsDNA viruses. Viruses. 16: 879.ArticlePubMedPMC
- Suraweera CD, Hinds MG, Kvansakul M. 2022. Structural insight into KsBcl-2 mediated apoptosis inhibition by Kaposi sarcoma associated herpes virus. Viruses. 14: 738.ArticlePubMedPMC
- Tidona CA, Darai G. 1997a. Molecular anatomy of lymphocystis disease virus. Arch Virol Suppl. 13: 49–56. Article
- Tidona CA, Darai G. 1997b. The complete DNA sequence of lymphocystis disease virus. Virology. 230: 207–216. Article
- Zhang H, Sheng X, Tang X, Xing J, Chi H, et al. 2023. Transcriptome analysis reveals molecular mechanisms of lymphocystis formation caused by lymphocystis disease virus infection in flounder (Paralichthys olivaceus). Front Immunol. 14: 1268851.ArticlePubMedPMC
- Zhang QY, Xiao F, Xie J, Li ZQ, Gui JF. 2004. Complete genome sequence of lymphocystis disease virus isolated from China. J Virol. 78: 6982–6994. ArticlePubMedPMCLink
References
Supplementary Information
References
Citations

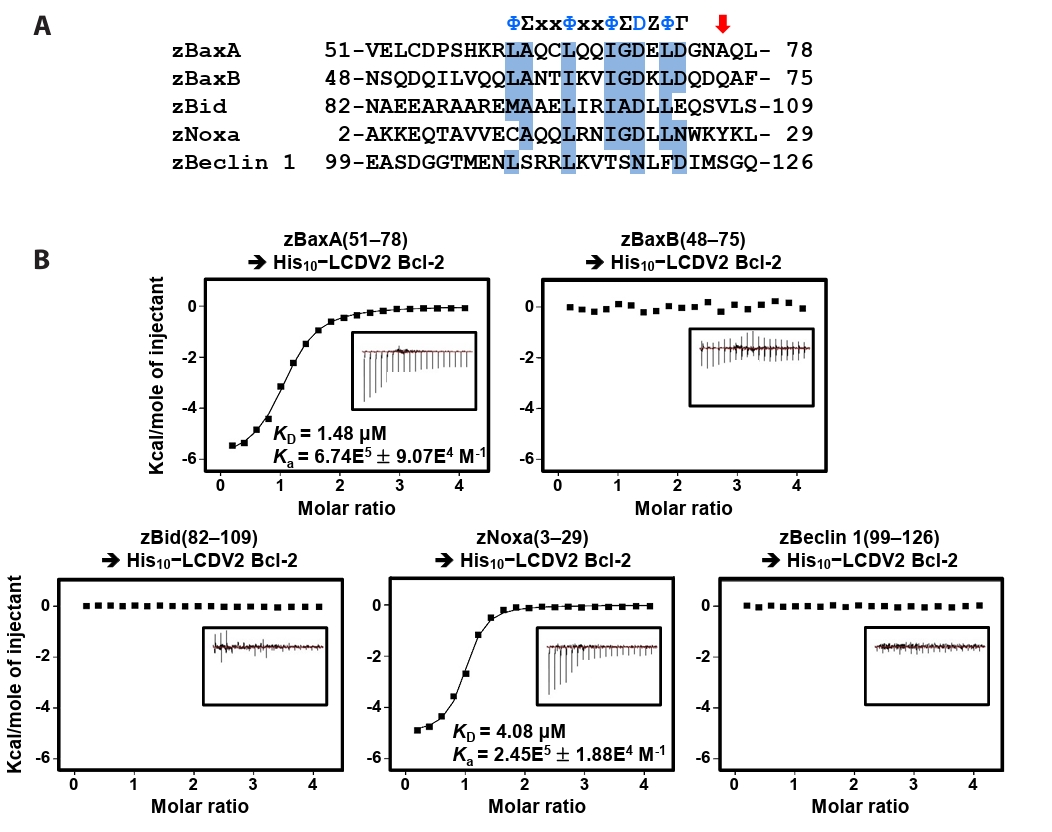
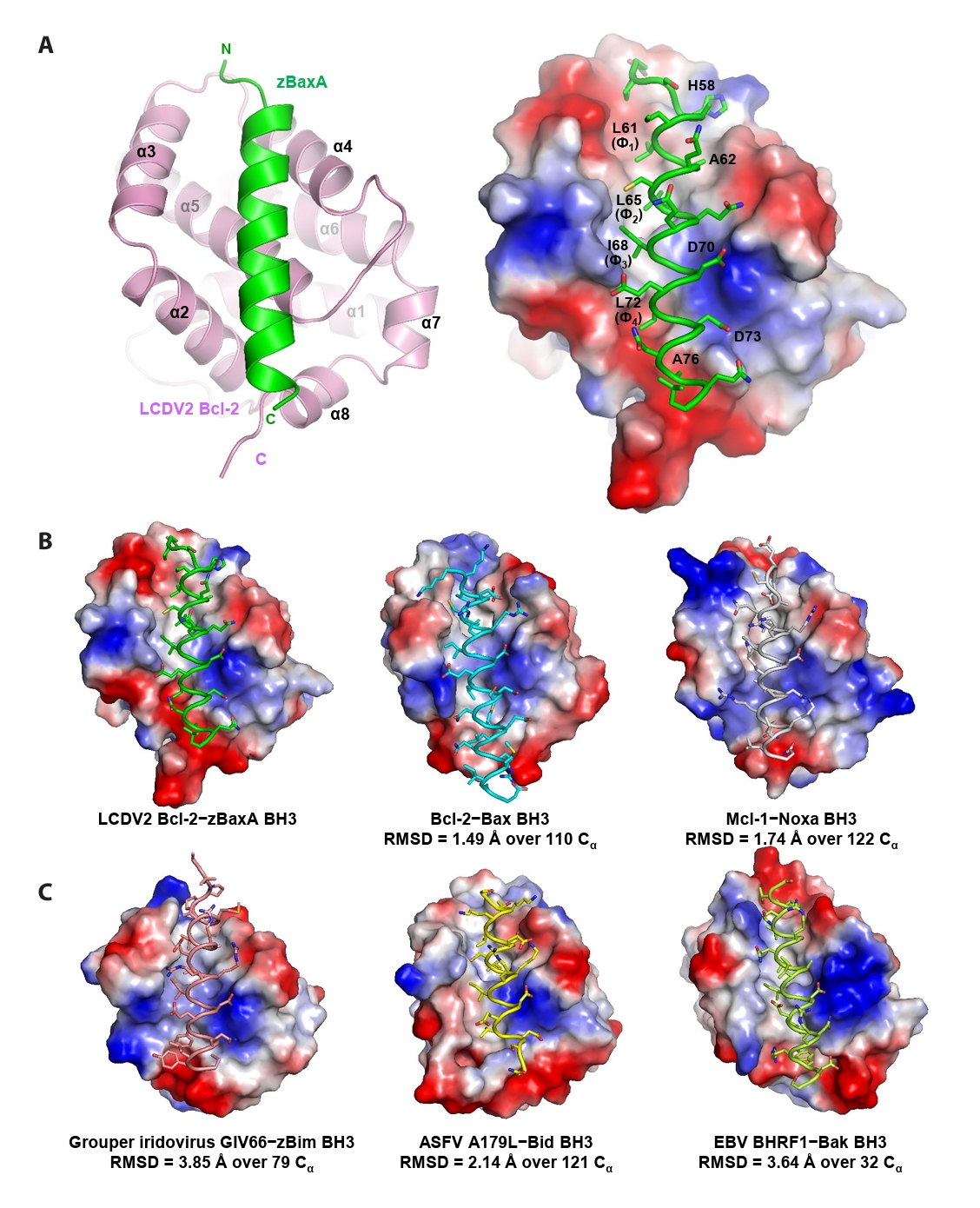
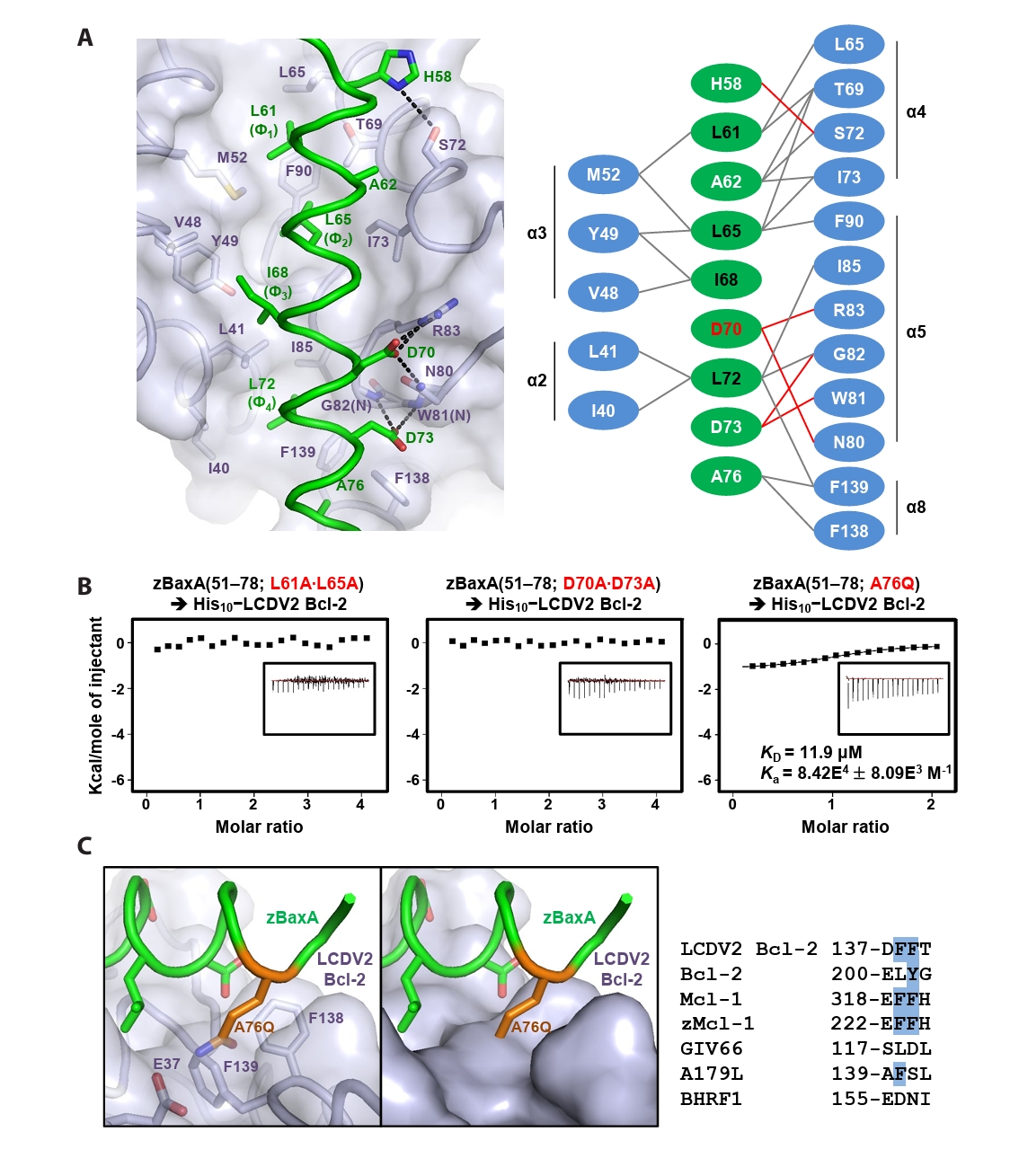
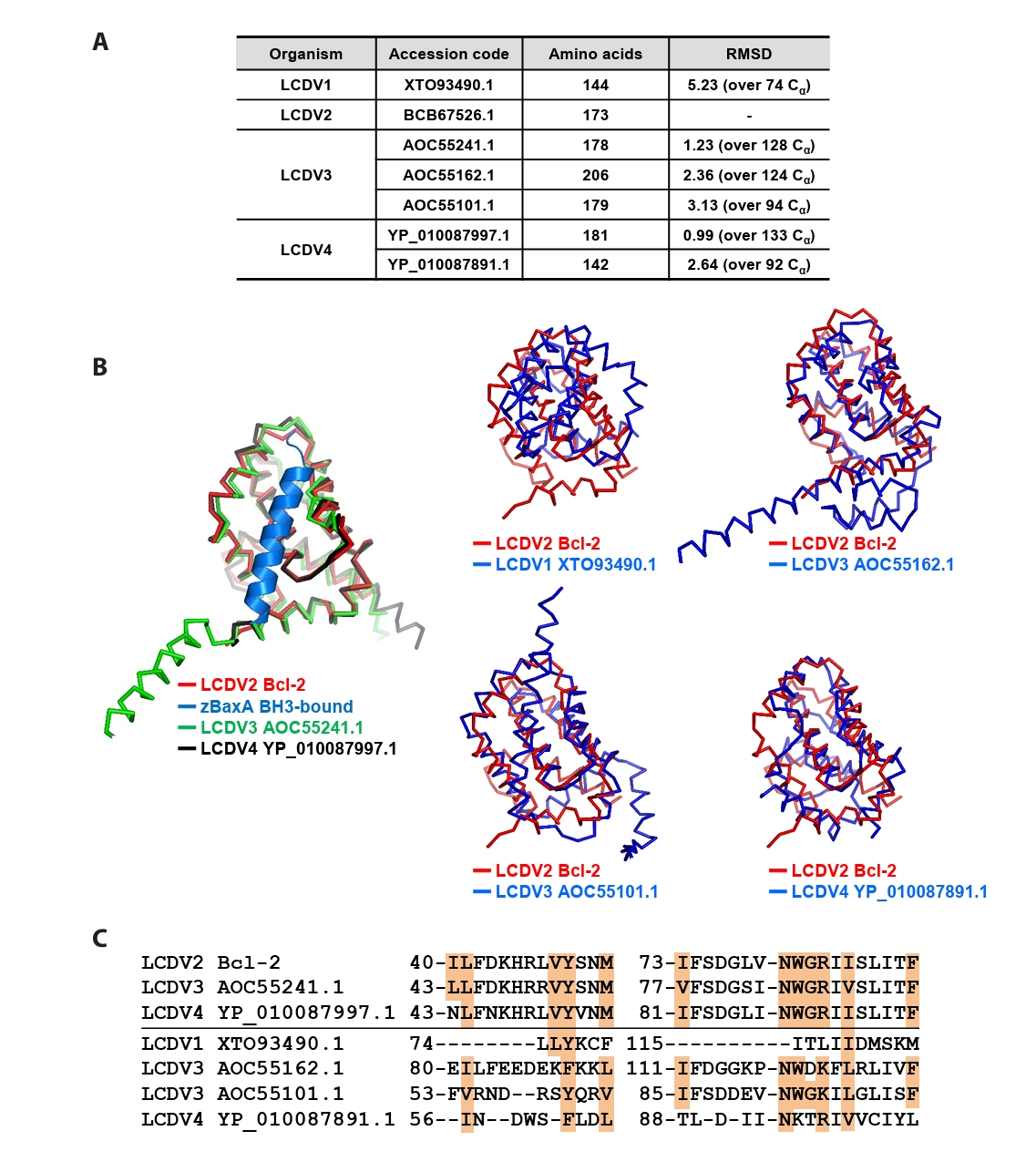
Fig. 1.
Fig. 2.
Fig. 3.
Fig. 4.
| Protein | LCDV2 Bcl-2−zBaxA(51–78) |
|---|---|
| Data collection | |
| Space group | P41212 |
| Unit cell dimensions | |
| a, b, c (Å) / a = β = γ (°) | 89.14, 89.14, 50.60 / 90 |
| Resolution (Å) | 50.0−2.3 (2.34−2.30) |
| Rsym |
11.6 (30.1) |
| I / σ (I) | 26.3 (3.9) |
| Completeness (%) | 96.8 (94.6) |
| Redundancy | 5.5 |
| Refinement | |
| Resolution (Å) | 50.0−2.3 |
| Number of reflections | 9304 |
| Rwork |
19.3 / 24.3 |
| Number of atoms | |
| Protein | 1133 |
| Peptide | 176 |
| Water | 28 |
| RMSD | |
| Bond lengths (Å) | 0.006 |
| Bond angles (°) | 0.728 |
| Ramachandran plot (%) | |
| Most favored region | 96.9 |
| Additionally allowed region | 3.1 |
| Average B-values (Å2) | |
| Protein | 43.5 |
| Peptide | 46.5 |
| Water | 45.6 |
The numbers in parentheses are statistics from the shell with the highest resolution.
Table 1.
TOP
 MSK
MSK
 ePub Link
ePub Link Cite this Article
Cite this Article