- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- For Contributors
- Policies
- E-Submission

- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- Policies
- For Contributors
Articles
- Page Path
- HOME > J. Microbiol > Ahead of print > Article
-
Review
High yield strategies for triterpenoid biosynthesis in cell factories - Mingzhu Zheng1,2,†, Chuang Liu1,2,†, Ceyuan Liu1,2, Jing Xie2, Gen Pan2, Can Zhong2,*, Jian Jin1,2,*
-
DOI: https://doi.org/10.71150/jm.2509018
Published online: April 21, 2026
1Hunan Academy of Chinese Medicine, Hunan University of Chinese Medicine, Changsha 410036, P. R. China
2Institute of Chinese Medicine Resources, Hunan Academy of Chinese Medicine, Changsha 410013, P. R. China
- *Correspondence Can Zhong canzhong651@163.com Jian Jin jinjian2016@163.com
- †These authors contributed equally to this work.
© The Microbiological Society of Korea
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
- 301 Views
- 11 Download
ABSTRACT
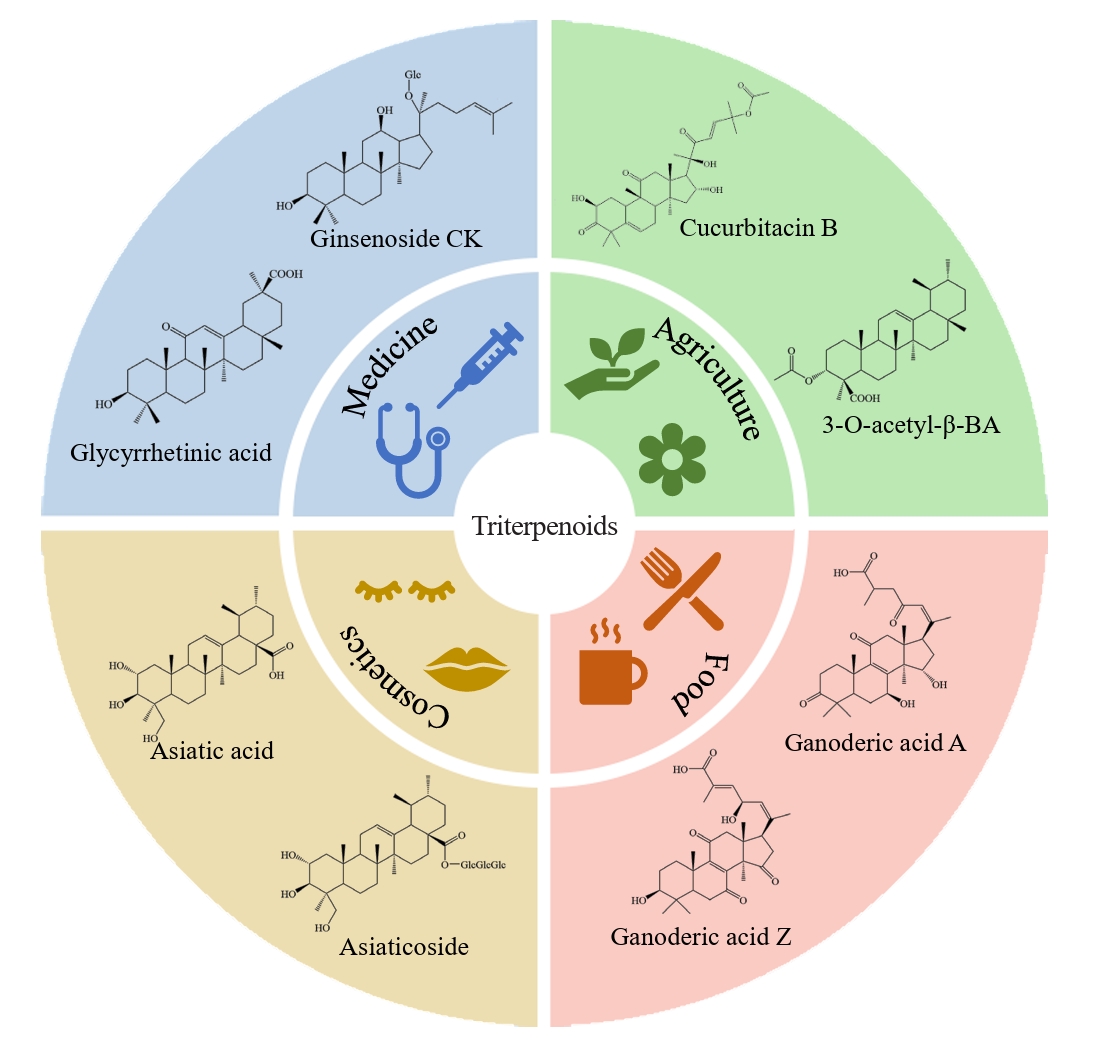
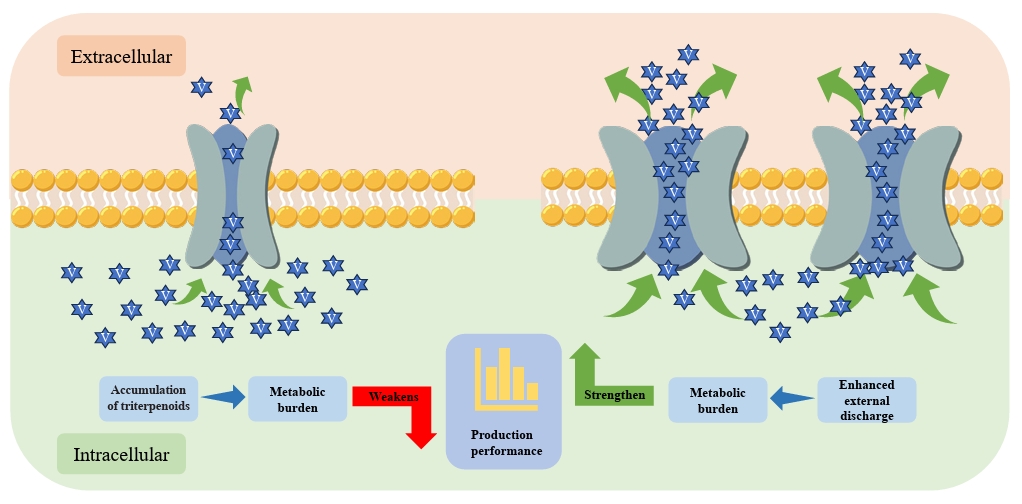
- Triterpenoids are natural products widely found in the plant kingdom and have various pharmacological effects such as anti-inflammatory, antioxidant and anti-tumour. However, the content of triterpenoids in medicinal plants is low, and it is difficult to purify and isolate them due to their complex structure. The efficient production of some triterpenoids in chassis organisms has been achieved by constructing a heterologous triterpenoid synthesis pathway in engineered strains such as yeast, modifying the key enzymes in the pathway, and adjusting the metabolism of yeast. Modification of key enzymes in the synthetic pathway is currently an effective strategy to enhance the heterologous synthesis of triterpenoids. This paper reviews the current research progress on the modification of key enzymes downstream in the synthetic pathway and the design of key enzymes around them to enhance triterpenoid production in five main areas: 1) increasing the supply of triterpenoid precursors; 2) inhibition of the natural sterol pathway; 3) fusion expression of related enzymes; 4) compartmentalisation of the metabolic pathway; and 5) tapping and enhancing the triterpenoid efflux pump. Finally, recent advances and applications of artificial intelligence (AI) in enzyme engineering and pathway design for triterpenoid biosynthesis are highlighted. Challenges and perspectives for further increasing the yield of triterpenoid synthesis in Saccharomyces cerevisiae are presented.
Introduction
Biosynthetic Pathways of Triterpenoids
Strategies for Improving Triterpenoid Biosynthesis
1. AI-assisted enzyme engineering for triterpenoid biosynthesis
2. AI in metabolic pathway design and optimization
Challenge and Prospect of Triterpenoid Production by Yeast
Acknowledgments
This work was supported by National Natural Science Foundation of China (No. 82474039, No. 82404805), Science and Technology Innovation Program of Hunan Province (No. 2025RC3227), Training Program for Excellent Young Innovators of Changsha (kq2506026), and the First-Class Discipline of Pharmaceutical Science of Hunan.
Conflict of Interest
The authors declare no conflicts of interest.
Supplementary Information
Fig. S1.
Fig. S2.
Fig. S3.
| Triterpenoid | Chassis organism | Key strategy | Production method | Titer | Reference |
|---|---|---|---|---|---|
| 7-Dehydrocholesterol | S. cerevisiae | Modular engineering, dynamic control | 5 L fed-batch fermentation | 2.87 g/L | Xiu et al. (2022) |
| Protopanaxadiol (PPD) | S. cerevisiae | Modular pathway engineering, metabolic flux optimization, Multi Strategy UGT engineering, computational simulation assistance | 10 L fed-batch fermentation | 11.02 g/L | Wang et al. (2019) |
| Ginsenoside Rh2 | S. cerevisiae | Modular pathway engineering, metabolic flux optimization, Multi Strategy UGT engineering, computational simulation assistance | 10 L fed-batch fermentation | 2.25 g/L | Wang et al. (2019) |
| Betulinic acid | S. cerevisiae | Metabolic engineering, compartment engineering, fermentation optimization | 5 L fed-batch fermentation | 205.74 mg/L | Tang et al. (2024) |
| α-Farnesene | Yarrowia lipolytica | Optimization of precursor supply, metabolic flow guidance, multi copy integration and fermentation process regulation | 1 L fed-batch fermentation | 25.55 g/L | Liu et al. (2019) |
| Squalene | S. cerevisiae | The whole squalene synthesis pathway (starting from acetyl coenzyme A) is reconstructed in peroxisome | 3 L fed-batch fermentation | 32.8 g/L | Ma et al. (2024) |
| Lycopene | S. cerevisiae | Multi module metabolic engineering, dynamic regulation of competitive pathways, cofactor optimization and efflux engineering | 50 ml shake flask fermentation | 343.7 mg/L | Huang et al. (2024) |
| Artemisinic acid | S. cerevisiae | Engineering transformation and optimization of yeast strains and innovation of fermentation process | Ethanol pulse feeding + IPM extraction fermentation | 25 g/L | Paddon et al. (2013) |
| No. | Strategy | Specific operation | Effect | Reference |
|---|---|---|---|---|
| 1 | Increase the supply of precursor substances | Replacement of the native ERG1 promoter with a strong promoter and substitution of endogenous ERG7 with a heterologous ERG7. | Increase the intracellular 2,3-oxidosqualene reserve. | Tang et al. (2024) |
| 2 | Utilization of a NADH-dependent HMG-CoA reductase (NADH-HMGR) from Silicobacterium malate to replace tHMG1. | Strengthened the MVA pathway activity, improving precursor supply for terpenoid biosynthesis. | Sun et al. (2024) | |
| 3 | Inhibition of the natural sterol pathway | Site-directed mutagenesis (F699T, I705K) introduced into ERG7. | Shift metabolic flux more towards downstream triterpenoid synthesis rather than sterol pathway. | Guo et al. (2022) |
| 4 | Regulating metabolic flux | Knockout of the transcription factor SIP4 regulating gluconeogenesis and ICL1. | Redirected carbon flux from gluconeogenesis toward target product biosynthesis. | Sun et al. (2024) |
| 5 | Overexpression of enzymes in the pre-2,3-oxidosqualene pathway. | Elevated cellular sterol content, alleviating growth inhibition caused by sterol depletion and stabilizing membrane integrity. | Guo et al. (2022) | |
| 6 | Improve the catalytic activity of enzyme | Site-directed mutagenesis (F222Y) introduced into the catalytic domain of CYP450-Uni25647. | Enhanced substrate binding affinity and catalytic activity of CYP450-Uni25647, improving triterpenoid modification efficiency. | Sun et al. (2024) |
| 7 | Site-saturation mutagenesis targeting the non-catalytic MXXXXR motif of pentacyclic triterpenoid synthases (PTSs). | Improved catalytic turnover rate by optimizing enzyme conformational dynamics, increasing triterpenoid yield by 18%. | Guo et al. (2024a) | |
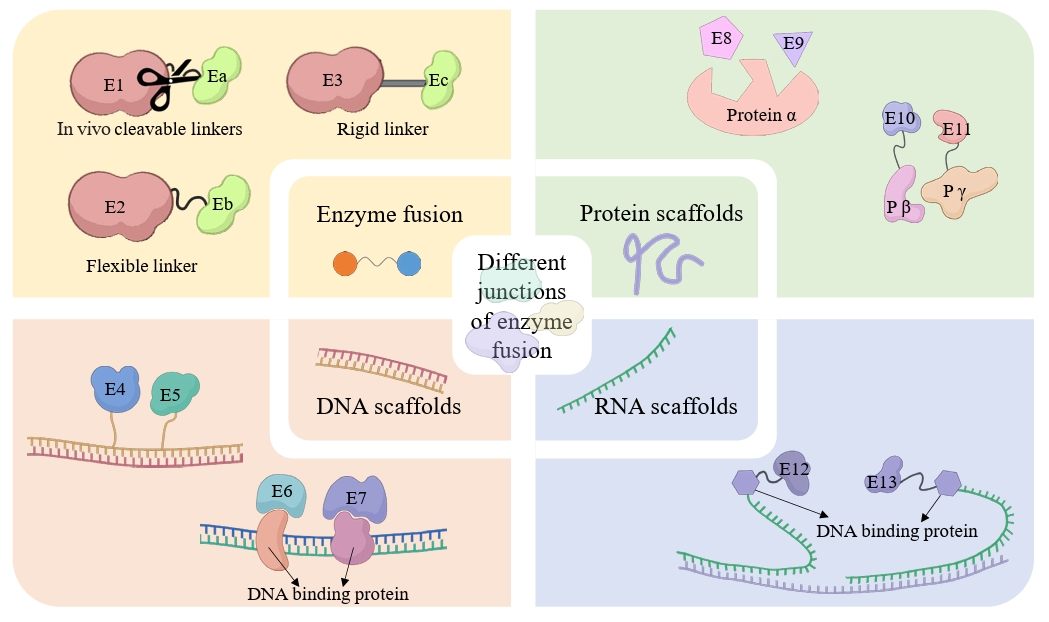
| 8 | Co-expression and fusion expression of related enzymes | Co-expression and fusion expression of CYP450 (BPLO) with cytochrome P450 reductase (CPR, ATR1). | Optimized redox coupling efficiency, reducing electron leakage and increasing hydroxylation activity by 37%. | Tang et al. (2024) |
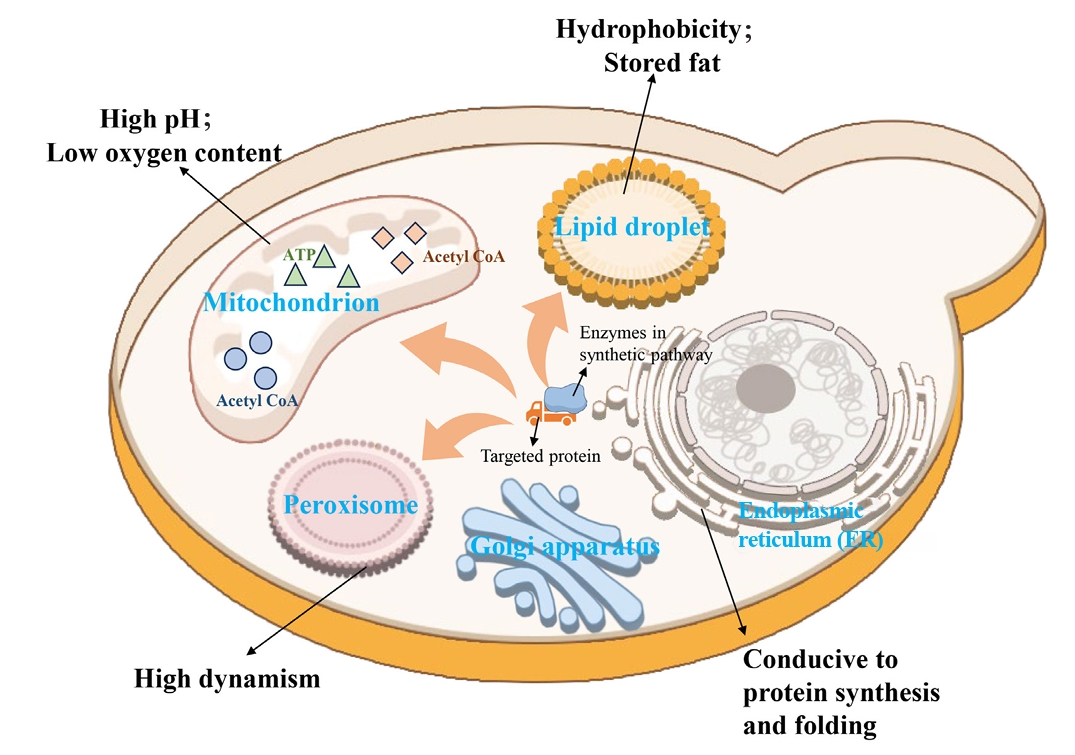
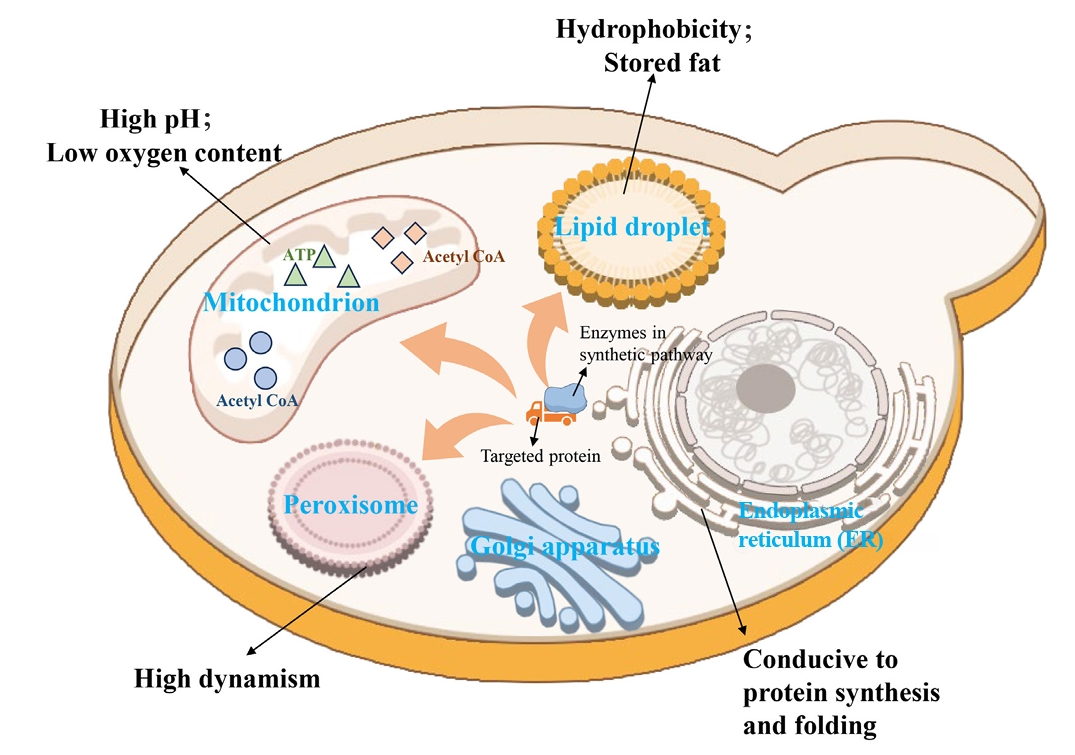
| 9 | Compartmentalization of metabolic pathways | Establishment of an orthogonal acetyl-CoA synthesis shortcut from CO2-derived acetate in peroxisomes. | Significantly enhance the accumulation of peroxisome squalene. | Ma et al. (2024) |
| 10 | Reconstruction of a complete squalene biosynthetic pathway from acetyl-CoA within peroxisomes. | Relieved the bottleneck of precursor supply and increased the titer of squalene by 4.5 times. | Ma et al. (2024) | |
| 11 | Co-localization of all enzymes in the squalene-to-betulinic acid pathway to lipid droplets. | The catalytic efficiency was improved by 62% through substrate channel effect. | Tang et al. (2024) |
- Aly SH, Elbadry AM, Doghish AS, El-Nashar HA. 2024. Unveiling the pharmacological potential of plant triterpenoids in breast cancer management: an updated review. Naunyn Schmiedebergs Arch Pharmacol. 397: 5571–5596. ArticlePubMedPMCPDF
- Bouin A, Zhang C, Lindley ND, Truan G, Lautier T. 2023. Exploring linker's sequence diversity to fuse carotene cyclase and hydroxylase for zeaxanthin biosynthesis. Metab Eng Commun. 16: e00222. ArticlePubMedPMC
- Bröker JN, Müller B, van Deenen N, Prüfer D, Schulze Gronover C. 2018. Upregulating the mevalonate pathway and repressing sterol synthesis in Saccharomyces cerevisiae enhances the production of triterpenes. Appl Microbiol Biotechnol. 102: 6923–6934. ArticlePubMedPMCPDF
- Bu X, Lin JY, Cheng J, Yang D, Duan CQ, et al. 2020. Engineering endogenous ABC transporter with improving ATP supply and membrane flexibility enhances the secretion of β-carotene in Saccharomyces cerevisiae. Biotechnol Biofuels. 13: 168.ArticlePubMedPMCPDF
- Chauhan S, Kerr A, Keogh B, Nolan S, Casey R, et al. 2021. An artificial-intelligence-discovered functional ingredient, NRT_N0G5IJ, derived from Pisum sativum, decreases HbA1c in a prediabetic population. Nutrients. 13: 1635.ArticlePubMedPMC
- da Silva LYS, Paulo CLR, Moura TF, Alves DS, Pessoa RT, et al. 2023. Antibacterial activity of the essential oil of Piper tuberculatum Jacq. fruits against multidrug-resistant strains: Inhibition of efflux pumps and β-lactamase. Plants. 12: 2377.ArticlePubMedPMC
- Dai Z, Liu Y, Zhang X, Shi M, Wang B, et al. 2013. Metabolic engineering of Saccharomyces cerevisiae for production of ginsenosides. Metab Eng. 20: 146–156. ArticlePubMed
- Du J, Yuan Y, Si T, Lian J, Zhao H. 2012. Customized optimization of metabolic pathways by combinatorial transcriptional engineering. Nucleic Acids Res. 40: e142. ArticlePubMedPMC
- Engle K, Kumar G. 2024. Tackling multi-drug resistant fungi by efflux pump inhibitors. Biochem Pharmacol. 226: 116400.ArticlePubMed
- Fan W, Huang Y, Zheng H, Li S, Li Z, et al. 2020. Ginsenosides for the treatment of metabolic syndrome and cardiovascular diseases: Pharmacology and mechanisms. Biomed Pharmacother. 132: 110915.ArticlePubMed
- Gao Y, Li J, Wang J, Li X, Li J, et al. 2020. Ginsenoside Rg1 prevent and treat inflammatory diseases: A review. Int Immunopharmacol. 87: 106805.ArticlePubMed
- Gómez-Bombarelli R, Wei JN, Duvenaud D, Hernández-Lobato JM, Sánchez-Lengeling B, et al. 2018. Automatic chemical design using a data-driven continuous representation of molecules. ACS Cent Sci. 4: 268–276. ArticlePubMedPMCLink
- Gorgulla C, Boeszoermenyi A, Wang ZF, Fischer PD, Coote PW, et al. 2020. An open-source drug discovery platform enables ultra-large virtual screens. Nature. 580: 663–668. ArticlePubMedPMCPDF
- Guo H, Chen T, Zhu H, Wang H, Huo YX. 2024a. Engineering amino acid residues of pentacyclic triterpene synthases for improving the activity. Appl Microbiol Biotechnol. 108: 195.ArticlePDF
- Guo H, Wang H, Chen T, Guo L, Blank LM, et al. 2022. Engineering critical amino acid residues of lanosterol synthase to improve the production of triterpenoids in Saccharomyces cerevisiae. ACS Synth Biol. 11: 2685–2696. ArticlePubMedLink
- Guo H, Wang H, Huo YX. 2020. Engineering critical enzymes and pathways for improved triterpenoid biosynthesis in yeast. ACS Synth Biol. 9: 2214–2227. ArticlePubMed
- Guo Q, Yang YX, Li DX, Ji XJ, Wu N, et al. 2024b. Advances in multi-enzyme co-localization strategies for the construction of microbial cell factory. Biotechnol Adv. 108453.Article
- Hakim SE, Liu S, Herzog R, Arafa A, de Vries J, et al. 2025. Expansion of the stereochemical space of triterpenes by mining noncanonical oxidosqualene cyclases across the diversity of green plants. J Am Chem Soc. 147: 10320–10330. ArticlePubMedPMCLink
- Hamberger B, Bak S. 2013. Plant P450s as versatile drivers for evolution of species-specific chemical diversity. Philos Trans R Soc B Biol Sci. 368: 20120426.ArticleLink
- Hannigan GD, Prihoda D, Palicka A, Soukup J, Klempir O, et al. 2019. A deep learning genome-mining strategy for biosynthetic gene cluster prediction. Nucleic Acids Res. 47: e110. ArticlePubMedPMCPDF
- Haringa C, Mudde RF, Noorman HJ. 2018. From industrial fermentor to CFD-guided downscaling: What have we learned? Biochem Eng J. 140: 57–71. Article
- Hoffmann MA, Nothias LF, Ludwig M, Fleischauer M, Gentry EC, et al. 2022. High-confidence structural annotation of metabolites absent from spectral libraries. Nat Biotechnol. 40: 411–421. ArticlePubMedPDF
- Hu J, Dong L, Outten CE. 2008. The redox environment in the mitochondrial intermembrane space is maintained separately from the cytosol and matrix. J Biol Chem. 283: 29126–29134. ArticlePubMedPMC
- Huang G, Li J, Lin J, Duan C, Yan G. 2024. Multi-modular metabolic engineering and efflux engineering for enhanced lycopene production in recombinant Saccharomyces cerevisiae. J Ind Microbiol Biotechnol. 51: kuae015.ArticlePubMedPMCPDF
- Huang Z, Zhang C, Xing XH. 2021. Chapter two - Design and construction of chimeric linker library with controllable flexibilities for precision protein engineering. In Merkx M. (ed.), Methods in enzymology, vol. 647, pp. 23–49. Academic Press. Article
- Jauch J. 2008. Total synthesis of azadirachtin—finally completed after 22 years. Angew Chem Int Ed. 47: 34–37. Article
- Jumper J, Evans R, Pritzel A, Green T, Figurnov M, et al. 2021. Highly accurate protein structure prediction with AlphaFold. Nature. 596: 583–589. ArticlePubMedPDF
- Koch M, Duigou T, Faulon JL. 2020. Reinforcement learning for bioretrosynthesis. ACS Synth Biol. 9: 157–168. ArticlePubMed
- Kretzschmar T, Burla B, Lee Y, Martinoia E, Nagy R. 2011. Functions of ABC transporters in plants. Essays Biochem. 50: 145–160. ArticlePubMedPDF
- Larroude M, Celinska E, Back A, Thomas S, Nicaud JM, et al. 2018. A synthetic biology approach to transform Yarrowia lipolytica into a competitive biotechnological producer of β-carotene. Biotechnol Bioeng. 115: 464–472. ArticlePubMedLink
- Lee JY, Kang CD, Lee SH, Park YK, Cho KM. 2015. Engineering cellular redox balance in Saccharomyces cerevisiae for improved production of L-lactic acid. Biotechnol Bioeng. 112: 751–758. ArticlePubMed
- Liu Y, Jiang X, Cui Z, Wang Z, Hou J. 2019. Engineering the oleaginous yeast Yarrowia lipolytica for production of α-farnesene. Biotechnol Biofuels. 12: 296.ArticlePubMedPMCPDF
- Liu Q, Liu Y, Li G, Savolainen O, Chen Y, et al. 2021. De novo biosynthesis of bioactive isoflavonoids by engineered yeast cell factories. Nat Commun. 12: 6085.ArticlePubMedPMCPDF
- Liu B, Ramsundar B, Kawthekar P, Shi J, Gomes J, et al. 2017. Retrosynthetic reaction prediction using neural sequence-to-sequence models. ACS Cent Sci. 3: 1103–1113. ArticlePubMedPMCLink
- Luo ZL, Zhang KL, Ma XJ, Guo YH. 2016. Research progress in synthetic biology of triterpen saponins. Chin Tradit Herb Drugs. 47: 1806–1814. Article
- Ma Y, Shang Y, Stephanopoulos G. 2024. Engineering peroxisomal biosynthetic pathways for maximization of triterpene production in Yarrowia lipolytica. Proc Natl Acad Sci USA. 121: e2314798121. ArticlePubMedPMC
- Mainali P, Khongsay N, Au-Yeung BSY, Zhang C, Selvarajoo K, et al. 2026. Kinetic modeling of terpenoid production in E. coli: insights into subpopulation emergence and process optimization. Microb Cell Fact. 25: 30.ArticlePDF
- Meadows AL, Hawkins KM, Tsegaye Y, Antipov E, Kim Y, et al. 2016. Rewriting yeast central carbon metabolism for industrial isoprenoid production. Nature. 537: 694–697. ArticlePubMedPDF
- Meijer D, Beniddir MA, Coley CW, Mejri YM, Öztürk M, et al. 2025. Empowering natural product science with AI: Leveraging multimodal data and knowledge graphs. Nat Prod Rep. 42: 654–662. ArticlePubMedPMC
- Mellor J, Grigoras I, Carbonell P, Faulon JL. 2016. Semisupervised Gaussian process for automated enzyme search. ACS Synth Biol. 5: 518–528. ArticlePubMed
- Mikulak-Klucznik B, Gołębiowska P, Bayly AA, Popik O, Klucznik T, et al. 2020. Computational planning of the synthesis of complex natural products. Nature. 588: 83–88. ArticlePubMedPDF
- Mo C, Bard M. 2005. A systematic study of yeast sterol biosynthetic protein-protein interactions using the split-ubiquitin system. Biochim Biophys Acta Mol Cell Biol Lipids. 1737: 152–160. Article
- Mo C, Milla P, Athenstaedt K, Ott R, Balliano G, et al. 2003. In yeast sterol biosynthesis the 3-keto reductase protein (Erg27p) is required for oxidosqualene cyclase (Erg7p) activity. Biochim Biophys Acta Mol Cell Biol Lipids. 1633: 68–74. Article
- Mühlenhoff U, Lill R. 2000. Biogenesis of iron-sulfur proteins in eukaryotes: a novel task of mitochondria that is inherited from bacteria. Biochim Biophys Acta Bioenerg. 1459: 370–382. Article
- Nielsen J. 2014. Synthetic biology for engineering acetyl coenzyme A metabolism in yeast. mBio. 5: e02153-14.ArticlePubMedPMCLink
- Paddon CJ, Westfall PJ, Pitera DJ, Benjamin K, Fisher K, et al. 2013. High-level semi-synthetic production of the potent antimalarial artemisinin. Nature. 496: 528–532. ArticlePubMed
- Peng B, Nielsen LK, Kampranis SC, Vickers CE. 2018. Engineered protein degradation of farnesyl pyrophosphate synthase is an effective regulatory mechanism to increase monoterpene production in Saccharomyces cerevisiae. Metab Eng. 47: 83–93. ArticlePubMed
- Pereira R, Mohamed ET, Radi MS, Herrgård MJ, Feist AM, et al. 2020. Elucidating aromatic acid tolerance at low pH in Saccharomyces cerevisiae using adaptive laboratory evolution. Proc Natl Acad Sci USA. 117: 27954–27961. ArticlePubMedPMC
- Pothoulakis G, Nguyen MT, Andersen ES. 2022. Utilizing RNA origami scaffolds in Saccharomyces cerevisiae for dCas9-mediated transcriptional control. Nucleic Acids Res. 50: 7176–7187. ArticlePubMedPMCPDF
- Repecka D, Jauniskis V, Karpus L, Rembeza E, Rokaitis I, et al. 2021. Expanding functional protein sequence spaces using generative adversarial networks. Nat Mach Intell. 3: 324–333. ArticlePDF
- Romero PA, Krause A, Arnold FH. 2013. Navigating the protein fitness landscape with Gaussian processes. Proc Natl Acad Sci USA. 110: E193–E201. ArticlePubMed
- Ryu JY, Kim HU, Lee SY. 2019. Deep learning enables high-quality and high-throughput prediction of enzyme commission numbers. Proc Natl Acad Sci USA. 116: 13996–14001. ArticlePubMedPMC
- Sarhene M, Ni JY, Duncan ES, Liu Z, Li S, et al. 2021. Ginsenosides for cardiovascular diseases; update on pre-clinical and clinical evidence, pharmacological effects and the mechanisms of action. Pharmacol Res. 166: 105481.ArticlePubMed
- Segler MH, Preuss M, Waller MP. 2018. Planning chemical syntheses with deep neural networks and symbolic AI. Nature. 555: 604–610. ArticlePubMedPDF
- Shen CR, Lan EI, Dekishima Y, Baez A, Cho KM, et al. 2011. Driving forces enable high-titer anaerobic 1-butanol synthesis in Escherichia coli. Appl Environ Microbiol. 77: 2905–2915. ArticlePubMedPMCLink
- Stephenson MJ, Owen C, Reed J, Osbourn A. 2025. Large-scale mining of plant genomes unlocks the diversity of oxidosqualene cyclases. Nat Chem Biol. 22: 249–259. ArticlePubMedPMCPDF
- Stokes JM, Yang K, Swanson K, Jin W, Cubillos-Ruiz A, et al. 2020. A deep learning approach to antibiotic discovery. Cell. 180: 688–702. ArticlePubMedPMC
- Struble TJ, Alvarez JC, Brown SP, Chytil M, Cisar J, et al. 2020. Current and future roles of artificial intelligence in medicinal chemistry synthesis. J Med Chem. 63: 8667–8682. ArticlePubMedPMCLink
- Sun C, Theodoropoulos C, Scrutton NS. 2019. Techno-economic assessment of microbial limonene production. Bioresour Technol. 300: 122666.ArticlePubMedPMC
- Sun W, Wan S, Liu C, Wang R, Zhang H, et al. 2024. Establishing cell suitability for high-level production of licorice triterpenoids in yeast. Acta Pharm Sin B. 14: 4134–4148. ArticlePubMedPMC
- Sun M, Ye Y, Xiao L, Duan X, Zhang Y, et al. 2017. Anticancer effects of ginsenoside Rg3. Int J Mol Med. 39: 507–518. ArticlePubMed
- Tang S, Ji W, Zhao Y, Zhang J, Wei D, et al. 2024. De novo biosynthesis of betulinic acid in engineered Saccharomyces cerevisiae. Bioorg Chem. 152: 107737.ArticlePubMed
- Tong Y, Luo YF, Gao W. 2021. Biosynthesis of paclitaxel using synthetic biology. Phytochem Rev. 21: 863–877. ArticlePDF
- Wang P, Wei W, Ye W, Li X, Zhao W, et al. 2019. Synthesizing ginsenoside Rh2 in Saccharomyces cerevisiae cell factory at high-efficiency. Cell Discov. 5: 5.ArticlePubMedPMCPDF
- Wu G, Yan Q, Jones JA, Tang YJ, Fong SS, et al. 2016. Metabolic burden: cornerstones in synthetic biology and metabolic engineering applications. Trends Biotechnol. 34: 652–664. ArticlePubMed
- Xiu X, Sun Y, Wu Y, Jin K, Qu L, et al. 2022. Modular remodeling of sterol metabolism for overproduction of 7-dehydrocholesterol in engineered yeast. Bioresour Technol. 360: 127572.ArticlePubMed
- Xu Y, Wang X, Zhang C, Zhou X, Xu X, et al. 2022. De novo biosynthesis of rubusoside and rebaudiosides in engineered yeasts. Nat Commun. 13: 3040.ArticlePubMedPMCPDF
- Yang KK, Wu Z, Arnold FH. 2019. Machine-learning-guided directed evolution for protein engineering. Nat Methods. 16: 687–694. ArticlePubMedPDF
- Yim H, Haselbeck R, Niu W, Pujol-Baxley C, Burgard A, et al. 2011. Metabolic engineering of Escherichia coli for direct production of 1,4-butanediol. Nat Chem Biol. 7: 445–452. ArticlePubMedPDF
- Zhang L, Tian X, Sun L, Mi K, Wang R, et al. 2024. Bacterial efflux pump inhibitors reduce antibiotic resistance. Pharmaceutics. 16: 170.ArticlePubMedPMC
- Zhao F, Bai P, Nan W, Li D, Zhang C, et al. 2019. A modular engineering strategy for high-level production of protopanaxadiol from ethanol by Saccharomyces cerevisiae. AIChE J. 65: 866–874. ArticleLink
- Zhou P, Xie W, He S, Sun Y, Meng X, et al. 2019. Ginsenoside Rb1 as an anti-diabetic agent and its underlying mechanism analysis. Cells. 8: 204.ArticlePubMedPMC
References
Supplementary Information
References
Citations

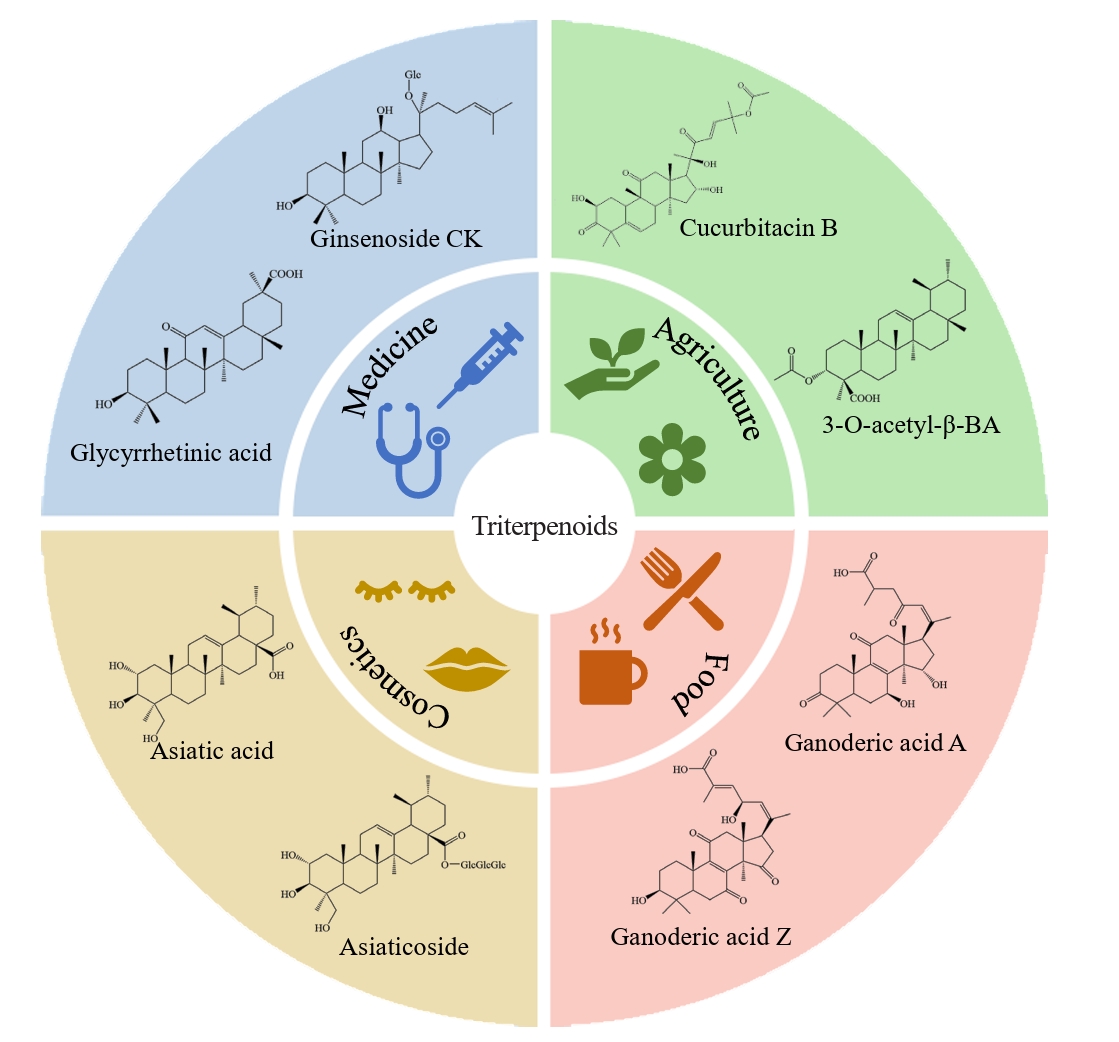
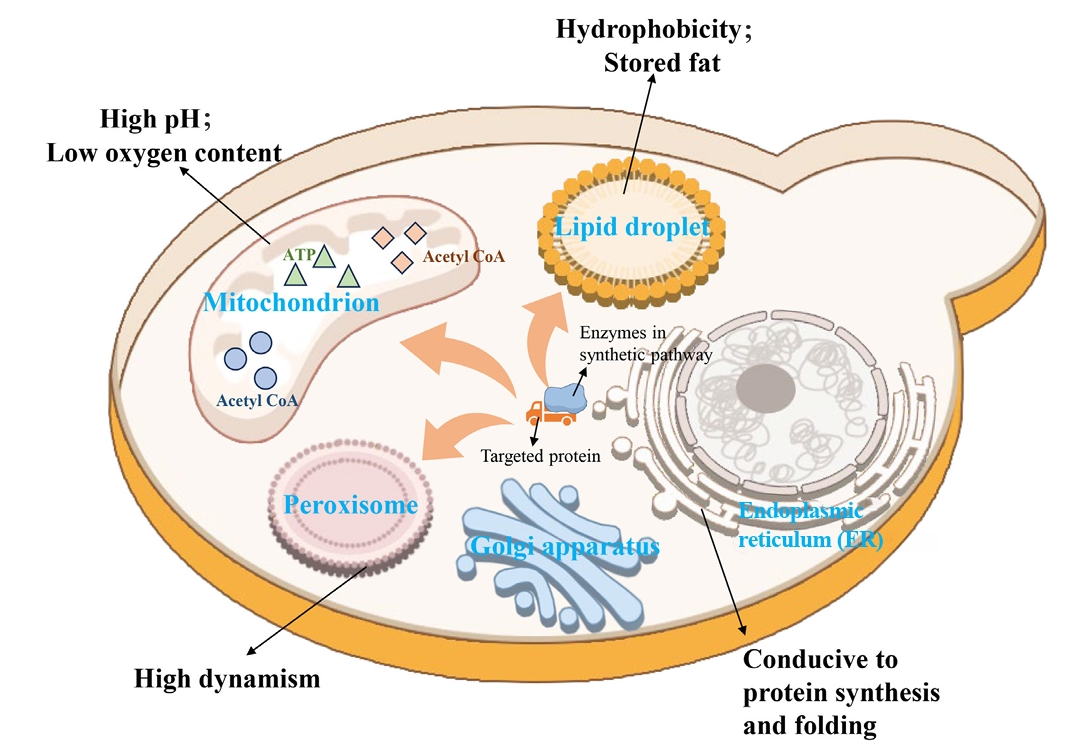
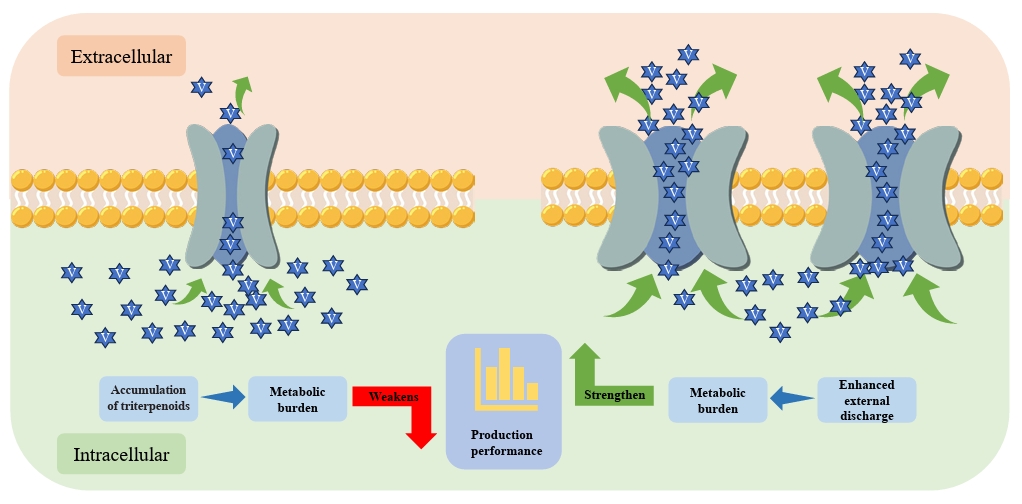
Fig. 1.
Fig. 2.
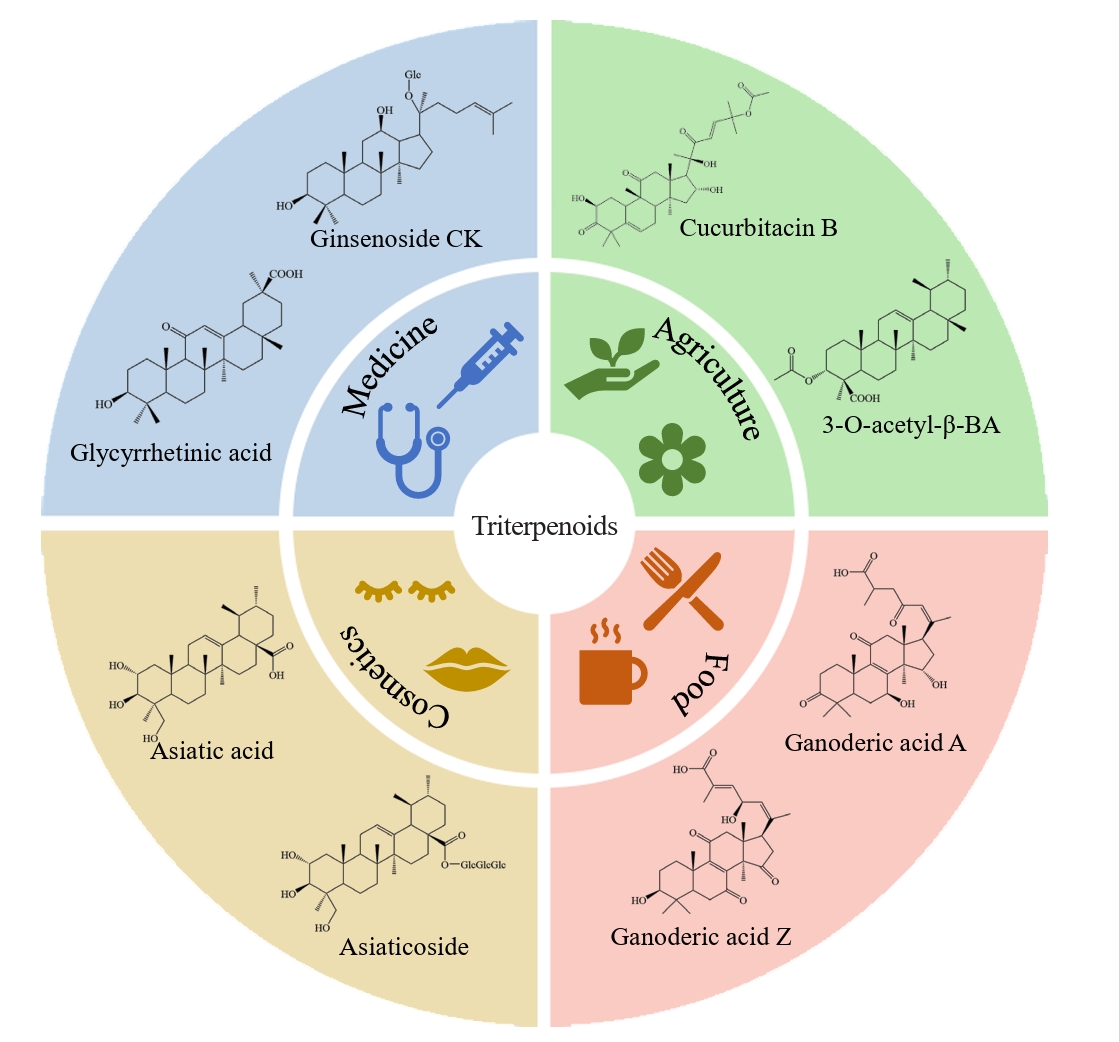
Fig. 3.
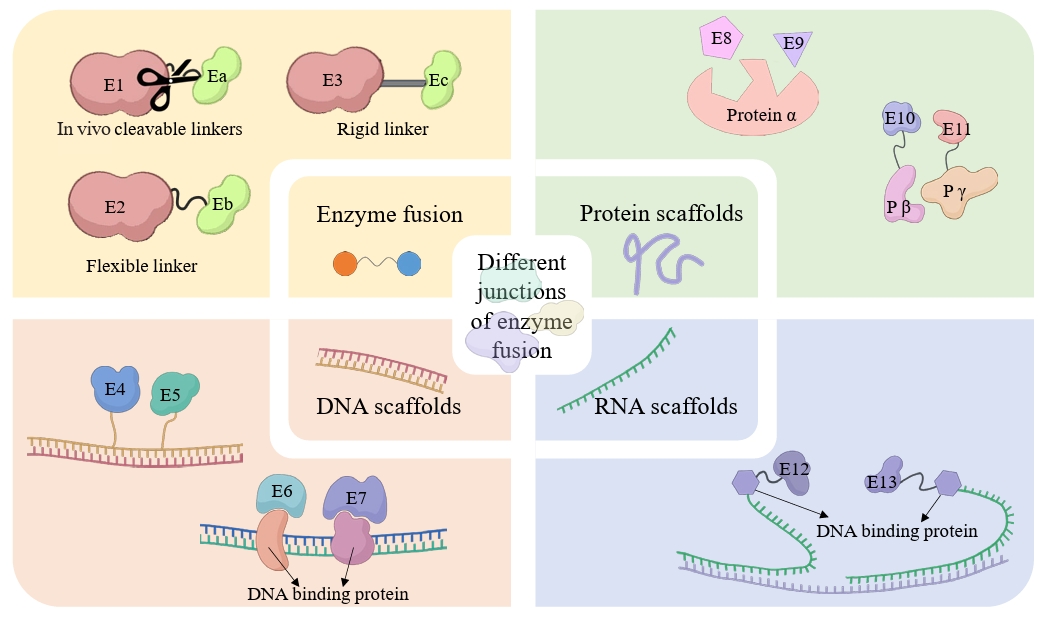
Fig. 4.
| Triterpenoid | Chassis organism | Key strategy | Production method | Titer | Reference |
|---|---|---|---|---|---|
| 7-Dehydrocholesterol | S. cerevisiae | Modular engineering, dynamic control | 5 L fed-batch fermentation | 2.87 g/L | |
| Protopanaxadiol (PPD) | S. cerevisiae | Modular pathway engineering, metabolic flux optimization, Multi Strategy UGT engineering, computational simulation assistance | 10 L fed-batch fermentation | 11.02 g/L | |
| Ginsenoside Rh2 | S. cerevisiae | Modular pathway engineering, metabolic flux optimization, Multi Strategy UGT engineering, computational simulation assistance | 10 L fed-batch fermentation | 2.25 g/L | |
| Betulinic acid | S. cerevisiae | Metabolic engineering, compartment engineering, fermentation optimization | 5 L fed-batch fermentation | 205.74 mg/L | |
| α-Farnesene | Yarrowia lipolytica | Optimization of precursor supply, metabolic flow guidance, multi copy integration and fermentation process regulation | 1 L fed-batch fermentation | 25.55 g/L | |
| Squalene | S. cerevisiae | The whole squalene synthesis pathway (starting from acetyl coenzyme A) is reconstructed in peroxisome | 3 L fed-batch fermentation | 32.8 g/L | |
| Lycopene | S. cerevisiae | Multi module metabolic engineering, dynamic regulation of competitive pathways, cofactor optimization and efflux engineering | 50 ml shake flask fermentation | 343.7 mg/L | |
| Artemisinic acid | S. cerevisiae | Engineering transformation and optimization of yeast strains and innovation of fermentation process | Ethanol pulse feeding + IPM extraction fermentation | 25 g/L |
| No. | Strategy | Specific operation | Effect | Reference |
|---|---|---|---|---|
| 1 | Increase the supply of precursor substances | Replacement of the native ERG1 promoter with a strong promoter and substitution of endogenous ERG7 with a heterologous ERG7. | Increase the intracellular 2,3-oxidosqualene reserve. | |
| 2 | Utilization of a NADH-dependent HMG-CoA reductase (NADH-HMGR) from Silicobacterium malate to replace tHMG1. | Strengthened the MVA pathway activity, improving precursor supply for terpenoid biosynthesis. | ||
| 3 | Inhibition of the natural sterol pathway | Site-directed mutagenesis (F699T, I705K) introduced into ERG7. | Shift metabolic flux more towards downstream triterpenoid synthesis rather than sterol pathway. | |
| 4 | Regulating metabolic flux | Knockout of the transcription factor SIP4 regulating gluconeogenesis and ICL1. | Redirected carbon flux from gluconeogenesis toward target product biosynthesis. | |
| 5 | Overexpression of enzymes in the pre-2,3-oxidosqualene pathway. | Elevated cellular sterol content, alleviating growth inhibition caused by sterol depletion and stabilizing membrane integrity. | ||
| 6 | Improve the catalytic activity of enzyme | Site-directed mutagenesis (F222Y) introduced into the catalytic domain of CYP450-Uni25647. | Enhanced substrate binding affinity and catalytic activity of CYP450-Uni25647, improving triterpenoid modification efficiency. | |
| 7 | Site-saturation mutagenesis targeting the non-catalytic MXXXXR motif of pentacyclic triterpenoid synthases (PTSs). | Improved catalytic turnover rate by optimizing enzyme conformational dynamics, increasing triterpenoid yield by 18%. | ||
| 8 | Co-expression and fusion expression of related enzymes | Co-expression and fusion expression of CYP450 (BPLO) with cytochrome P450 reductase (CPR, ATR1). | Optimized redox coupling efficiency, reducing electron leakage and increasing hydroxylation activity by 37%. | |
| 9 | Compartmentalization of metabolic pathways | Establishment of an orthogonal acetyl-CoA synthesis shortcut from CO2-derived acetate in peroxisomes. | Significantly enhance the accumulation of peroxisome squalene. | |
| 10 | Reconstruction of a complete squalene biosynthetic pathway from acetyl-CoA within peroxisomes. | Relieved the bottleneck of precursor supply and increased the titer of squalene by 4.5 times. | ||
| 11 | Co-localization of all enzymes in the squalene-to-betulinic acid pathway to lipid droplets. | The catalytic efficiency was improved by 62% through substrate channel effect. |
Table 1.
Table 2.
TOP
 MSK
MSK
 ePub Link
ePub Link Cite this Article
Cite this Article