- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- For Contributors
- Policies
- E-Submission

- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- Policies
- For Contributors
Articles
- Page Path
- HOME > J. Microbiol > Ahead of print > Article
-
Review
I53-50: Engineered icosahedral protein cage for modular vaccine nanoplatform - Ke Liang, Shuang Wu, Sihang Dong, Tao Xu*, Hongtao Wang*
-
DOI: https://doi.org/10.71150/jm.2511020
Published online: April 6, 2026
Anhui Province Key Laboratory of Immunology in Chronic Diseases, Laboratory Medicine Experimental Center, Laboratory Medicine College, Bengbu Medical University, Bengbu 233030, P. R. China
- *Correspondence Tao Xu taoxu@bbmu.edu.cn Hongtao Wang hongtaowang@bbmu.edu.cn
© The Microbiological Society of Korea
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
- 42 Views
- 6 Download
ABSTRACT
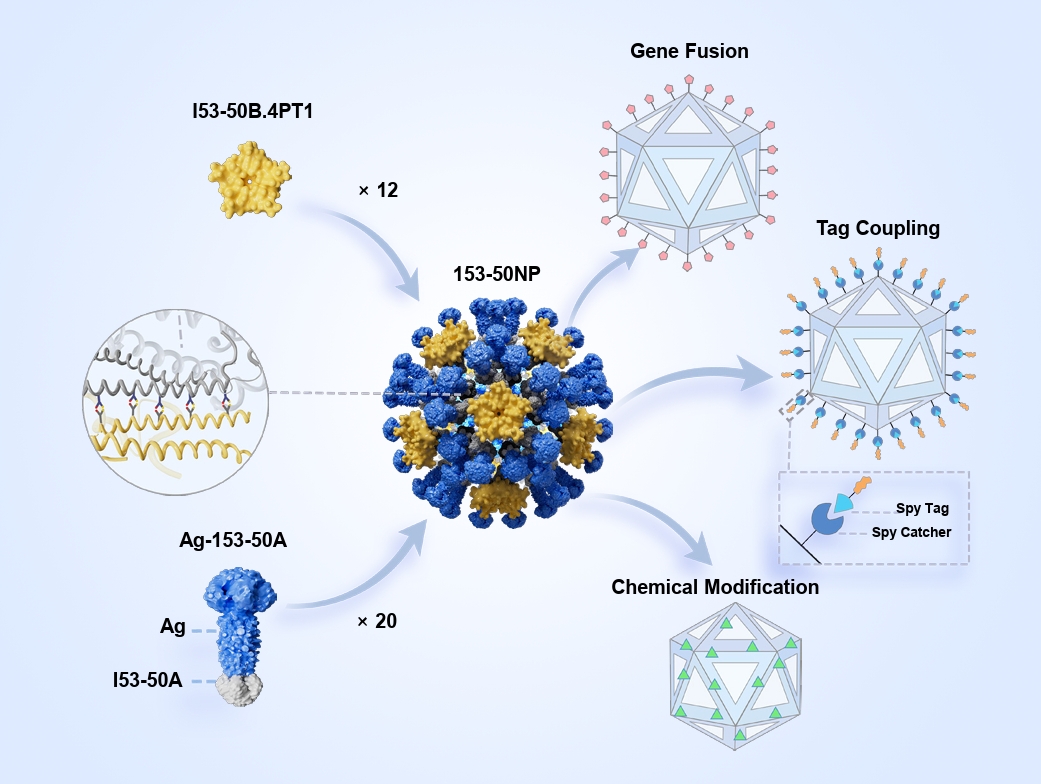
- I53-50 is a computationally designed, self-assembling protein nanoparticle (NP) that forms a stable icosahedral structure composed of 120 protein subunits coordinated through precise interfacial interactions. Through unique intelligent regulation, I53-50 exhibits sensitivity to environmental signals and display multimodal “nano-smart” properties. I53-50 has a variety of modifiable surface-active sites, which facilitates precise chemical modification, gene fusion, tag coupling, and other functionalizations, thereby promoting effective lymphatic uptake and optimizing the immune response. I53-50 NPs show great potential in vaccine development, drug delivery, and biomaterials, representing a model fusion of computational biology and nanomedicine and offering a versatile tool for precision medicine.
Structure and Properties of I53-50 NPs
Properties of I53-50
Biomedical Applications of I53-50 NPs
Discussion
Acknowledgments
The associate editors thank the authors, and all the reviewers and editors who undoubtedly contributed to the realization of this Research Topic.
Conflicts of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Funding
This work was supported by Natural Science Foundation of the Anhui Higher Education Institutions (2024AH051295), Bengbu City science and technology project (2023hm01), Key Projects of Natural Science Foundation of Bengbu Medical University (2023byzd020).
| Functionalization strategy | Key characteristics | Advantages | Limitations |
|---|---|---|---|
| Chemical modification | Targeting intrinsic active residues (preferentially N-terminal/C-terminal regions) of I53-50 subunits to conjugate functional groups or molecules via chemical reactions, thereby achieving protein functional modification and enhancement (Brouwer et al., 2019). | 1. Site-specificity: Enables precise targeting of defined amino acid sites. | 1. Potential structural perturbation: Chemical reactions may affect the native protein conformation and stability. |
| 2. High flexibility: Compatible with a wide range of chemical reagents and reactions. | 2. Risk of heterogeneity: Conjugation reactions can yield heterogeneous products. | ||
| 3. Overcomes spatial constraints: Linker design allows fine-tuning of the distance between the antigen and the nanoparticle surface (Sletten and Bertozzi, 2009). | 3. Dependence on reactive residues: Efficiency is limited by the accessibility of reactive amino acids (Sletten and Bertozzi, 2009; Stephanopoulos and Francis, 2011). | ||
| Tag coupling | Introducing specific tags into I53-50 subunits via genetic engineering or chemical methods, and achieving site-specific covalent coupling through tag-specific recognition for antigen display, purification, or targeting (Bruun et al., 2018). | 1. Stable covalent linkage: Antigen is covalently and stably integrated into the assembly. | 1. Insert size limitation: Large antigen inserts may impair subunit folding and self-assembly. |
| 2. High-density & Uniform display: Enables precise, ordered presentation of 60-240 antigen copies per particle. | 2. Design complexity: Requires optimization of linkers (flexible/rigid) to balance antigen presentation and structural integrity. | ||
| 3. Streamlined production: Single-step expression and self-assembly simplify manufacturing (Zakeri et al., 2012). | 3. Long redesign cycles: New antigens require de novo gene construction and validation (Veggiani et al., 2016). | ||
| Gene fusion | Fusing the coding gene of the target antigen with the modified I53-50 subunit gene via recombinant DNA technology, allowing the fusion protein to self-assemble into functional nanoparticles with antigen display (Marcandalli et al., 2019). | 1. Modularity & Versatility: "Plug-and-play" platform for rapid interchange of different SpyTag-fused antigens. | 1. Requires tag engineering: Both carrier and antigen require genetic modification, which may affect their native properties. |
| 2. Site-specific & Efficient: Forms irreversible isopeptide bonds under mild conditions, ensuring high coupling efficiency and homogeneity. | 2. Potential tag immunogenicity: The tags themselves could elicit immune responses (typically low). | ||
| 3. Enhanced stability: Covalent linkage improves complex stability during storage and in vivo (Chen et al., 2013). | 3. Additional conjugation step: Involves an in vitro coupling step compared to genetic fusion (Bird et al., 1988). |
| Parameter / Feature | I53-50NP | VLP | Ferritin | Mi3 |
|---|---|---|---|---|
| Subunit composition | Composed of 24 trimeric (I53-50A) and 12 pentameric (I53-50B) subunits assembling with strict T = 1 icosahedral symmetry (Bale et al., 2016) | Typically assembles from identical subunits via quasi-equivalent T = 3 or T = 4 icosahedral symmetry, leading to potential structural variability (Fuenmayor et al., 2017) | 24 subunits self-assemble into a cage with octahedral (O) symmetry (Zhang et al., 2021) | Trimeric subunits assemble into 2D/3D lattices or finite particles with D3 (dihedral) symmetry (Eom et al., 2024) |
| Thermal stability (Tm) | Exceptionally high. Melting temperature (Tm) > 95°C (for variant I53-50A.1NT1) and remains intact after 1-h incubation at 70°C (Walls et al., 2020) | Moderate to high. Varies by source; e.g., HBcAg VLP Tm ~70–75°C, but may exhibit kinetic instability (Mohsen and Bachmann, 2022) | High. Human heavy-chain ferritin (HFH) Tm ~75°C, but stability varies across species and is pH-sensitive (Zhang et al., 2021) | High. Designed Mi3 protein exhibits Tm ~77°C (Liu et al., 2021) |
| Assembly mechanism | Strictly controlled co-assembly. A and B components must mix in a precise stoichiometric ratio to form uniform ~50 nm particles. Enables mosaic display of distinct antigens (Wargacki et al., 2021) | Condition-dependent self-assembly. Spontaneous assembly under specific conditions (pH, ionic strength) can lead to heterogeneity in size and morphology (Nooraei et al., 2021) | pH-dependent assembly/Disassembly. Assembles at neutral/alkaline pH and disassembles at acidic pH. Useful for cargo loading but may raise concerns about in vivo stability in certain microenvironments (Chen et al., 2022) | Self-assembly into extended arrays or finite particles. Achieving uniform, finite-sized nanoparticles requires precise design control (Wu et al., 2025) |
| Functionalization | Highly flexible & Precise. Supports genetic fusion (to N/C-termini or specific loops of A/B subunits), chemical conjugation, and tag coupling. Allow precise presentation of 60 or 120 antigen copies with controlled orientation (Wargacki et al., 2021) | Flexible but potentially disruptive. Common genetic fusion to N/C-termini or the major immunodominant (Mohsen and Bachmann, 2022) region (MIR). Insertions can interfere with assembly, and copy number and orientation control is less precise (Mohsen and Bachmann, 2022) | Primarily N/C-terminal genetic fusion. Due to octahedral symmetry, maximum valency is limited to 24 copies, resulting in lower antigen density (Munir et al., 2024) | Primarily genetic fusion to specific sites on the trimeric subunit. Its dihedral symmetry results in antigen geometry distinct from icosahedral presentation (Wu et al., 2025) |
| Name of the vaccine | Construction methods | Immunization effects | Type of connection | Assembly efficiency and conditions |
|---|---|---|---|---|
| RBD-I53-50NP (Kang et al., 2021) | ΔN1-SpyCatcher first binds to I53-50A1.1PT1 and I53-50B.4PT1, and then fuses with SpyTag attached to the C-terminal end of the SARS-CoV-2 RBD, resulting in ΔN1-SpyCatcher-I53-50NP | RBD-I53-50NP showed stronger affinity for the receptor ACE2 and the neutralizing antibody CB6, indicating good affinity for BCR and triggering higher titers of antibody | tag coupling | I53-50A1.1PT1 and I53-50B.4PT1 were incubated in vitro at 250 mM NaCl, 50 mM Tris-HCl (pH 8.0), containing 5% (v/v) glycerol, and purified by size exclusion chromatography (SEC). No free RBD antigen or residual I53-50 dimer subunits were detected, with high subunit assembly and antigen loading conversion rates. |
| SARS-CoV2 S-I53-50NP (Arunachalam et al., 2021; Kang et al., 2021) | SARS-CoV-2 S-I53-50NP formed by fusing the C-terminus of the pre-fused SARS-CoV-2 S protein to the N-terminus of the I53-50A variant of I53-50A.1NT1, followed by incubation with I53-50B.4PT1 | Enhanced activation of SARS-CoV-2 S protein-specific B cells in vitro induces a strong NAb response | gene fusion | SARS-CoV-2 S-I53-50A.1NT1 and I53-50B.4PT1 were mixed in buffer at 4°C or room temperature for several hours under the following conditions: 50 mM Tris (pH 8), 150 mM NaCl, 100 mM L-arginine, and 5% (w/v) sucrose. Assembly efficiency is typically very high (> 90%), yielding monodisperse nanoparticles with highly uniform morphology. |
| RBD-8G/12G/16G-I53-50NP (Arunachalam et al., 2021) | SARS-CoV-2 RBD was genetically fused to the N-terminus of I53-50A via a linker of 8, 12, and 16 glycine and serine residues, and later fused to I53-50B.4PT1 to form RBD-I53-50NP | RBD-NP immunization induced a robust and long-lasting neutralizing antibody response, eliciting a strong antigen-specific CD4 T-cell response that skillfully neutralized the mutant strain | gene fusion | I53-50A and I53-50B4.PT1 were incubated at room temperature (25–30°C) for 1–2 h in a buffer containing 250 mM NaCl, 50 mM Tris-HCl pH 7.4–8.0, and 5% glycerol. The subunit assembly conversion rate approached 100% (size exclusion chromatography (SEC) showed no free subunit peaks), with a polydispersity index (PDI) < 0.1 (determined by dynamic light scattering (DLS)). |
| E2E1-I53-50NP (Sliepen et al., 2022) | The N-terminus of I53-50A1.NT1 is fused to the C-terminus of E2E1, which later fuses with I53-50B.4PT1 to form E2E1-I53-50NP | The number of neutralizing viruses in animals immunized with E2E1-NPs was significantly higher than that in animals receiving monoimmunization, and both the width and potency of Nab were significantly improved, with an increase in potency of up to 80-fold | gene fusion | E2E1-I53-50A and I53-50B.4PT1 were incubated overnight at 4°C in Tris-buffered saline (TBS, pH ~7.5) supplemented with 5% glycerol or L-arginine, achieving assembly efficiencies typically > 90%. Both SEC monomodal profiles and NS-EM images indicate the product consists of highly uniform protein nanoparticles, with no detectable free protein or aggregates. |
| p67C-I53-50NP (Lacasta et al., 2023) | ECF p67C is linked to the I53-50A N-terminus and the p67C-I53-50A fusion gene formed is fused to I53-50B.4PT1 to form p67c-I53-50NP | p67C-I53-50 immunized animals produced high p67C-specific IgG1 and IgG2 antibody titers as well as strong CD4+ T-cell responses, with the highest amount of IFN-γ secretion | gene fusion | p67C‑I53‑50A and I53-50B.4PT1 were incubated at room temperature (RT) for 30 min to 2 h in 50 mM Tris‑HCl, pH 8.0, 250–500 mM NaCl, supplemented with 50 mM glycine or 0.75% CHAPS, at room temperature (RT) for 30 min to 2 h. Size exclusion chromatography (SEC) revealed a single symmetric peak for the assembled nanoparticles, indicating a homogeneous product. Dynamic light scattering (DLS) showed a uniform particle size distribution consistent with the expected dimensions. |
| gB-I53-50-NP (Sun et al., 2023) | HCV viral gB gene fused to the N-terminus of I53-50A1 via C-terminal fusion, forming gB-I53-50A1 fused to I53-50B.4PT1 to form gB-I53-50NP | Induction of serum polyclonal antibodies by gB-I53-50NP vaccine in monkeys significantly protects humanized mice against EBV infection and lymphomagenesis, and the protective effect persists for a long period of time up to 10 weeks after immunization of monkeys | gene fusion | I53-50A1 and I53-50B4.PT1 can be efficiently assembled by incubating them for 1 h in assembly buffer (250 mM NaCl, 50 mM Tris pH 8.0, 5% glycerol). |
| SOSIP-I53-50NP (Brinkkemper et al., 2024) | HIV viral SOSIP Env trimer fused to I53-50A to form the SOSIP-I53-50A construct, which later fused to I53-50B.4PT1 to form SOSIP-I53-50NP | SOSIP-I53-50 NP in immunized animals, binds antibodies with reactive IgG titers up to 50–100 μg/ml, neutralizes viruses with titers ID50 > 100, Abs targets a wide range of neutralizing epitopes. | gene fusion | SOSIP-I53-50A and I53-50B.4PT1 were incubated overnight at 4°C in 25 mM Tris-HCl (pH 8.0), 500 mM NaCl, 5% glycerol at 4°C overnight. Dynamic light scattering analysis revealed the hydrodynamic radius of the assembled nanoparticles ranged from 246 to 292 Å with a low polydispersity index, indicating uniform and monodisperse particle size distribution. |
| DS-Cav1-I53-50 and Sc9-10-I53-50 (Hu et al., 2025) | The extracellular domain sequences of DS-Cav1 or Sc9-10 were fused with the I53-50A subunit gene to construct the DS-Cav1-I53-50A and Sc9-10-I53-50A fusion protein. The fusion protein was secreted and expressed in a stable Chinese hamster ovary (CHO) cell line, while the I53-50B subunit was expressed in Escherichia coli BL21 (DE3). The two subunits self-assembled in a 3:1 mass ratio to form DS-Cav1-I53-50 and Sc9-10-I53-50 nanoparticles. | In mouse models, total IgG titers, D25 competitive neutralization antibody titers, and live virus neutralization antibody titers against RSV A2 strain were elevated. Due to the antigenic properties of Sc9-10, Sc9-10-I53-50 exhibited slightly higher immunogenicity than DS-Cav1-I53-50. | gene fusion | DS-Cav1-I53-50A and Sc9-10-I53-50A were mixed with I53-50B in a buffer system of 50 mM Tris-HCl, 300 mM NaCl, 0.75% CHAPS, pH 7.4, and incubated at room temperature (RT) for 2 h. Purified by size exclusion chromatography (SEC), the particles exhibited uniform size and high assembly efficiency. |
| DS2-I53-50 (Jiang et al., 2025) | The DS2 antigen gene was directly fused with the I53-50A gene to construct the DS2-I53-50A fusion protein. The DS2-I53-50A fusion protein was transiently expressed in Expi293F mammalian cells, while the I53-50B subunit was induced in Escherichia coli BL21(DE3). Following separate purifications, the two subunits self-assembled in a 1:1 molar ratio to form stable DS2-I53-50 nanoparticles. | Following immunization of BALB/c mice, the induced DS2-specific IgG titer was 2.9 times that of free DS2, with the IgG1/IgG2a ratio decreasing to 2.6, indicating a Th1-biased response. This significantly promoted the proliferation of germinal center B cells (CD19⁺B220⁺CD95⁺GL-7⁺) and follicular helper T cells (CD4⁺CXCR5⁺PD-1⁺), activated dendritic cells and macrophages, and expanded the CD4⁺ central/ effector memory T cell populations. | gene fusion | Mix DS2-I53-50A with I53-50B in PBS (containing 5% glycerol) and incubate at 25°C for 3 h. Size exclusion chromatography shows that all particles assembled successfully, with the most uniform chromatographic peak observed for I53-50. |
- Almalik A, Benabdelkamel H, Masood A, Alanazi IO, Alradwan I, et al. 2017. Hyaluronic acid coated chitosan nanoparticles reduced the immunogenicity of the formed protein corona. Sci Rep. 7: 10542.ArticlePubMedPMCPDF
- Arunachalam PS, Walls AC, Golden N, Atyeo C, Fischinger S, et al. 2021. Adjuvanting a subunit COVID-19 vaccine to induce protective immunity. Nature. 594: 253–258. PubMed
- Bachmann MF, Jennings GT. 2010. Vaccine delivery: a matter of size, geometry, kinetics and molecular patterns. Nat Rev Immunol. 10: 787–796. ArticlePubMedPDF
- Bai Z, Cao Y, Liu W, Li J. 2021. The SARS-CoV-2 nucleocapsid protein and its role in viral structure, biological functions, and a potential target for drug or vaccine mitigation. Viruses. 13: 1115.ArticlePubMedPMC
- Bale JB, Gonen S, Liu Y, Sheffler W, Ellis D, et al. 2016. Accurate design of megadalton-scale two-component icosahedral protein complexes. Science. 353: 389–394. ArticlePubMedPMC
- Bale JB, Park RU, Liu Y, Gonen S, Gonen T, et al. 2015. Structure of a designed tetrahedral protein assembly variant engineered to have improved soluble expression. Protein Sci. 24: 1695–1701. ArticlePubMedPMCLink
- Becicka WM, Bielecki PA, Lorkowski ME, Moon TJ, Zhang Y, et al. 2021. The effect of PEGylation on the efficacy and uptake of an immunostimulatory nanoparticle in the tumor immune microenvironment. Nanoscale Adv. 3: 4961–4972. ArticlePubMedPMC
- Bhardwaj P, Bhatia E, Sharma S, Ahamad N, Banerjee R. 2020. Advancements in prophylactic and therapeutic nanovaccines. Acta Biomater. 108: 1–21. ArticlePubMedPMC
- Bird RE, Hardman KD, Jacobson JW, Johnson S, Kaufman BM, et al. 1988. Single-chain antigen-binding proteins. Science. 242: 423–426. ArticlePubMed
- Boyoglu-Barnum S, Ellis D, Gillespie RA, Hutchinson GB, Park Y, et al. 2021. Quadrivalent influenza nanoparticle vaccines induce broad protection. Nature. 592: 623–628. ArticlePubMedPMCPDF
- Brinkkemper M, Kerster G, Brouwer PJM, Tran AS, Torres JL, et al. 2024. Mosaic and mixed HIV-1 glycoprotein nanoparticles elicit antibody responses to broadly neutralizing epitopes. PLoS Pathog. 20: e1012558. ArticlePubMedPMC
- Brodin JD, Ambroggio XI, Tang C, Parent KN, Baker TS, et al. 2012. Metal-directed, chemically tunable assembly of one-, two- and three-dimensional crystalline protein arrays. Nat Chem. 4: 375–382. ArticlePubMedPMCPDF
- Brouwer PJM, Antanasijevic A, Berndsen Z, Yasmeen A, Fiala B, et al. 2019. Enhancing and shaping the immunogenicity of native-like HIV-1 envelope trimers with a two-component protein nanoparticle. Nat Commun. 10: 4272.ArticlePubMedPMCPDF
- Brouwer PJM, Brinkkemper M, Maisonnasse P, Dereuddre-Bosquet N, Grobben M, et al. 2021. Two-component spike nanoparticle vaccine protects macaques from SARS-CoV-2 infection. Cell. 184: 1188–1200. ArticlePubMedPMC
- Brouwer PJM, Sanders RW. 2019. Presentation of HIV-1 envelope glycoprotein trimers on diverse nanoparticle platforms. Curr Opin HIV AIDS. 14: 302–308. ArticlePubMedPMC
- Bruun TUJ, Andersson AC, Draper SJ, Howarth M. 2018. Engineering a rugged nanoscaffold to enhance plug-and-display vaccination. ACS Nano. 12: 8855–8866. ArticlePubMedPMCLink
- Buldun CM, Jean JX, Bedford MR, Howarth M. 2018. SnoopLigase catalyzes peptide-peptide locking and enables solid-phase conjugate isolation. J Am Chem Soc. 140: 3008–3018. ArticlePubMed
- Chen H, Tan X, Han X, Ma L, Dai H, et al. 2022. Ferritin nanocage based delivery vehicles: from single-, co- to compartmentalized-encapsulation of bioactive or nutraceutical compounds. Biotechnol Adv. 61: 108037.ArticlePubMed
- Chen H, Zhen Z, Todd T, Chu PK, Xie J. 2013. Nanoparticles for improving cancer diagnosis. Mater Sci Eng R Rep. 74: 35–69. ArticlePubMedPMC
- Dauparas J, Anishchenko I, Bennett N, Bai H, Ragotte RJ, et al. 2022. Robust deep learning-based protein sequence design using ProteinMPNN. Science. 378: 49–56. ArticlePubMedPMC
- Delehanty JB, Boeneman K, Bradburne CE, Robertson K, Bongard JE, et al. 2010. Peptides for specific intracellular delivery and targeting of nanoparticles: implications for developing nanoparticle-mediated drug delivery. Ther Deliv. 1: 411–433. ArticlePubMed
- Eom S, Jun H, Kim E, Min D, Kim H, et al. 2024. Developing porous protein cage nanoparticles as cargo-loadable and ligand-displayable modular delivery nanoplatforms. ACS Appl Mater Interfaces. 16: 58464–58476. ArticlePubMedLink
- Frank C, Khoshouei A, Fuβ L, Schiwietz D, Putz D, et al. 2024. Scalable protein design using optimization in a relaxed sequence space. Science. 386: 439–445. ArticlePubMedPMC
- Fuenmayor J, Gòdia F, Cervera L. 2017. Production of virus-like particles for vaccines. New Biotechnol. 39: 174–180. Article
- Goodsell DS, Olson AJ. 2000. Structural symmetry and protein function. Annu Rev Biophys Biomol Struct. 29: 105–153. ArticlePubMed
- Gou S, Geng W, Zou Y, Chen F, He T, et al. 2024. Glutathione-responsive and hydrogen sulfide self-generating nanocages based on self-weaving technology to optimize cancer immunotherapy. ACS Nano. 18: 9871–9885. ArticlePubMedLink
- Grueninger D, Treiber N, Ziegler MOP, Koetter JWA, Schulze M, et al. 2008. Designed protein-protein association. Science. 319: 206–209. ArticlePubMed
- He L, Tzarum N, Lin X, Shapero B, Sou C, et al. 2020. Proof of concept for rational design of hepatitis C virus E2 core nanoparticle vaccines. Sci Adv. 6: eaaz6225. ArticlePubMedPMC
- Hsia Y, Mout R, Sheffler W, Edman NI, Vulovic I, et al. 2021. Design of multi-scale protein complexes by hierarchical building block fusion. Nat Commun. 12: 2294.ArticlePubMedPMCPDF
- Hu Z, Tian S, Zhou Y, Wang Y, Li Y, et al. 2025. Nanoparticle vaccine based on the pre-fusion F glycoprotein of respiratory syncytial virus elicits robust protective immune responses. J Virol. 99: e00903-25. ArticlePubMedPMCLink
- Hura GL, Menon AL, Hammel M, Rambo RP, Poole FLN, et al. 2009. Robust, high-throughput solution structural analyses by small angle X-ray scattering (SAXS). Nat Methods. 6: 606–612. ArticlePubMedPMCPDF
- Janin J, Bahadur RP, Chakrabarti P. 2008. Protein-protein interaction and quaternary structure. Q Rev Biophys. 41: 133–180. ArticlePubMed
- Jia X, Zhao Q, Xiong Y. 2015. HIV suppression by host restriction factors and viral immune evasion. Curr Opin Struct Biol. 31: 106–114. ArticlePubMedPMC
- Jiang J, Hu H, Cao L, Mao N, Zhu Z, et al. 2025. Comparative evaluation of three nanoparticle vaccines targeting the prefusion F protein of respiratory syncytial virus: immunogenicity and protective efficacy. Int J Nanomedicine. 20: 9945–9961. PubMedPMC
- Kanekiyo M, Joyce MG, Gillespie RA, Gallagher JR, Andrews SF, et al. 2019. Mosaic nanoparticle display of diverse influenza virus hemagglutinins elicits broad B cell responses. Nat Immunol. 20: 362–372. ArticlePubMedPMCPDF
- Kang Y, Sun C, Zhuang Z, Yuan R, Zheng Q, et al. 2021. Rapid development of SARS-CoV-2 spike protein receptor-binding domain self-assembled nanoparticle vaccine candidates. ACS Nano. 15: 2738–2752. ArticlePubMedLink
- Keeble AH, Turkki P, Stokes S, Khairil Anuar INA, Rahikainen R, et al. 2019. Approaching infinite affinity through engineering of peptide-protein interaction. Proc Natl Acad Sci USA. 116: 26523–26533. ArticlePubMedPMC
- Kreuter J. 1996. Nanoparticles and microparticles for drug and vaccine delivery. J Anat. 189: 503–505. PubMedPMC
- Lacasta A, Kim HC, Kepl E, Gachogo R, Chege N, et al. 2023. Design and immunological evaluation of two-component protein nanoparticle vaccines for East Coast fever. Front Immunol. 13: 1015840. Article
- Lai Y, Reading E, Hura GL, Tsai K, Laganowsky A, et al. 2014. Structure of a designed protein cage that self-assembles into a highly porous cube. Nat Chem. 6: 1065–1071. ArticlePubMedPMCPDF
- Lawrence MC, Colman PM. 1993. Shape complementarity at protein/protein interfaces. J Mol Biol. 234: 946–950. ArticlePubMed
- Lee S, Kibler RD, Ahn G, Hsia Y, Borst AJ, et al. 2025. Four-component protein nanocages designed by programmed symmetry breaking. Nature. 638: 546–552. ArticlePubMedPDF
- Lin T, Chen Z, Usha R, Stauffacher CV, Dai JB, et al. 1999. The refined crystal structure of cowpea mosaic virus at 2.8 A resolution. Virology. 265: 20–34. ArticlePubMed
- Liu Z, Xu H, Han G, Tao L, Lu Y, et al. 2021. A self-assembling nanoparticle: implications for the development of thermostable vaccine candidates. Int J Biol Macromol. 183: 2162–2173. ArticlePubMed
- Maia A, Tarannum M, Lérias JR, Piccinelli S, Borrego LM, et al. 2024. Building a better defense: expanding and improving natural killer cells for adoptive cell therapy. Cells. 13: 451.ArticlePubMedPMC
- Mao C, Deng F, Zhu W, Xie L, Wang Y, et al. 2024. In situ editing of tumour cell membranes induces aggregation and capture of PD-L1 membrane proteins for enhanced cancer immunotherapy. Nat Commun. 15: 9723.ArticlePubMedPMCPDF
- Marcandalli J, Fiala B, Ols S, Perotti M, de van der Schueren W, et al. 2019. Induction of potent neutralizing antibody responses by a designed protein nanoparticle vaccine for respiratory syncytial virus. Cell. 176: 1420–1431. PubMedPMC
- Mohsen MO, Bachmann MF. 2022. Virus-like particle vaccinology, from bench to bedside. Cell Mol Immunol. 19: 993–1011. ArticlePubMedPMCPDF
- Munir I, Nazir F, Yesiloz G. 2024. Unlocking nature's potential: ferritin as a universal nanocarrier for amplified cancer therapy testing via 3D microtissues. ACS Appl Mater Interfaces. 16: 70187–70204. ArticlePubMedPMCLink
- Nooraei S, Bahrulolum H, Hoseini ZS, Katalani C, Hajizade A, et al. 2021. Virus-like particles: preparation, immunogenicity and their roles as nanovaccines and drug nanocarriers. J Nanobiotechnol. 19: 59.ArticlePDF
- Padilla JE, Colovos C, Yeates TO. 2001. Nanohedra: using symmetry to design self assembling protein cages, layers, crystals, and filaments. Proc Natl Acad Sci USA. 98: 2217–2221. ArticlePubMedPMC
- Pancera M, Zhou T, Druz A, Georgiev IS, Soto C, et al. 2014. Structure and immune recognition of trimeric pre-fusion HIV-1 Env. Nature. 514: 455–461. ArticlePubMedPMCPDF
- Pascha MN, Ballegeer M, Roelofs MC, Meuris L, Albulescu IC, et al. 2024. Nanoparticle display of neuraminidase elicits enhanced antibody responses and protection against influenza A virus challenge. NPJ Vaccines. 9: 97.ArticlePubMedPMCPDF
- Rahikainen R, Rijal P, Tan TK, Wu H, Andersson AC, et al. 2021. Overcoming symmetry mismatch in vaccine nanoassembly through spontaneous amidation. Angew Chem Int Ed Engl. 60: 321–330. ArticlePubMedLink
- Ramos-Pan L, Touzani A, Fernandez-Bertolez N, Fraga S, Laffon B, et al. 2024. Impact of gold nanoparticle exposure on genetic material. Mutat Res Genet Toxicol Environ Mutagen. 900: 503827.ArticlePubMed
- Reddy ST, van der Vlies AJ, Simeoni E, Angeli V, Randolph GJ, et al. 2007. Exploiting lymphatic transport and complement activation in nanoparticle vaccines. Nat Biotechnol. 25: 1159–1164. ArticlePubMedPDF
- Shen J, Ma M, Shafiq M, Yu H, Lan Z, et al. 2022. Microfluidics-assisted engineering of pH/enzyme dual-activatable ZIF@polymer nanosystem for co-delivery of proteins and chemotherapeutics with enhanced deep-tumor penetration. Angew Chem Int Ed Engl. 61: e202113703. ArticlePubMedLink
- Sletten EM, Bertozzi CR. 2009. Bioorthogonal chemistry: fishing for selectivity in a sea of functionality. Angew Chem Int Ed. 48: 6974–6998. Article
- Sliepen K, Radić L, Capella-Pujol J, Watanabe Y, Zon I, et al. 2022. Induction of cross-neutralizing antibodies by a permuted hepatitis C virus glycoprotein nanoparticle vaccine candidate. Nat Commun. 13: 7271.ArticlePubMedPMCPDF
- Song JY, Choi WS, Heo JY, Lee JS, Jung DS, et al. 2022. Safety and immunogenicity of a SARS-CoV-2 recombinant protein nanoparticle vaccine (GBP510) adjuvanted with AS03: A randomised, placebo-controlled, observer-blinded phase 1/2 trial. EClinicalMedicine. 51: 101569.ArticlePubMedPMC
- Stephanopoulos N, Francis MB. 2011. Choosing an effective protein bioconjugation strategy. Nat Chem Biol. 7: 876–884. ArticlePubMedPDF
- Sun C, Kang Y, Fang X, Liu Y, Bu G, et al. 2023. A gB nanoparticle vaccine elicits a protective neutralizing antibody response against EBV. Cell Host Microbe. 31: 1882–1897. ArticlePubMed
- Tang H, Chu W, Xiong J, Wu H, Cheng L, et al. 2024. Seeking cells, targeting bacteria: a cascade-targeting bacteria-responsive nanosystem for combating intracellular bacterial infections. Small. 20: e2311967. ArticlePubMed
- van Gils MJ, van den Kerkhof TLGM, Ozorowski G, Cottrell CA, Sok D, et al. 2016. An HIV-1 antibody from an elite neutralizer implicates the fusion peptide as a site of vulnerability. Nat Microbiol. 2: 16199.ArticlePubMedPMC
- Veggiani G, Nakamura T, Brenner MD, Gayet RV, Yan J, et al. 2016. Programmable polyproteams built using twin peptide superglues. Proc Natl Acad Sci USA. 113: 1202–1207. ArticlePubMedPMC
- Walls AC, Fiala B, Schäfer A, Wrenn S, Pham MN, et al. 2020. Elicitation of potent neutralizing antibody responses by designed protein nanoparticle vaccines for SARS-CoV-2. Cell. 183: 1367–1382.e17.ArticlePubMedPMC
- Wargacki AJ, Wörner TP, van de Waterbeemd M, Ellis D, Heck AJR, et al. 2021. Complete and cooperative in vitro assembly of computationally designed self-assembling protein nanomaterials. Nat Commun. 12: 883.ArticlePubMedPMCPDF
- Wu H, Weng R, Li J, Huang Z, Tie X, et al. 2025. Self-assembling protein nanoparticle platform for multivalent antigen delivery in vaccine development. Int J Pharm. 676: 125597.ArticlePubMed
- Yang Y, Dai M, Wilson TM, Omelchenko I, Klimek JE, et al. 2011. Na+/K+-ATPase α1 identified as an abundant protein in the blood-labyrinth barrier that plays an essential role in the barrier integrity. PLoS One. 6: e16547. ArticlePubMedPMC
- Yao N, Xiao W, Wang X, Marik J, Park SH, et al. 2009. Discovery of targeting ligands for breast cancer cells using the one-bead one-compound combinatorial method. J Med Chem. 52: 126–133. ArticlePubMedPMC
- Zakeri B, Fierer JO, Celik E, Chittock EC, Schwarz-Linek U, et al. 2012. Peptide tag forming a rapid covalent bond to a protein, through engineering a bacterial adhesin. Proc Natl Acad Sci USA. 109: E690–E697. ArticlePubMedPMC
- Zhang B, Tang G, He J, Yan X, Fan K. 2021. Ferritin nanocage: a promising and designable multi-module platform for constructing dynamic nanoassembly-based drug nanocarrier. Adv Drug Deliv Rev. 176: 113892.ArticlePubMed
References
Figure & Data
References
Citations

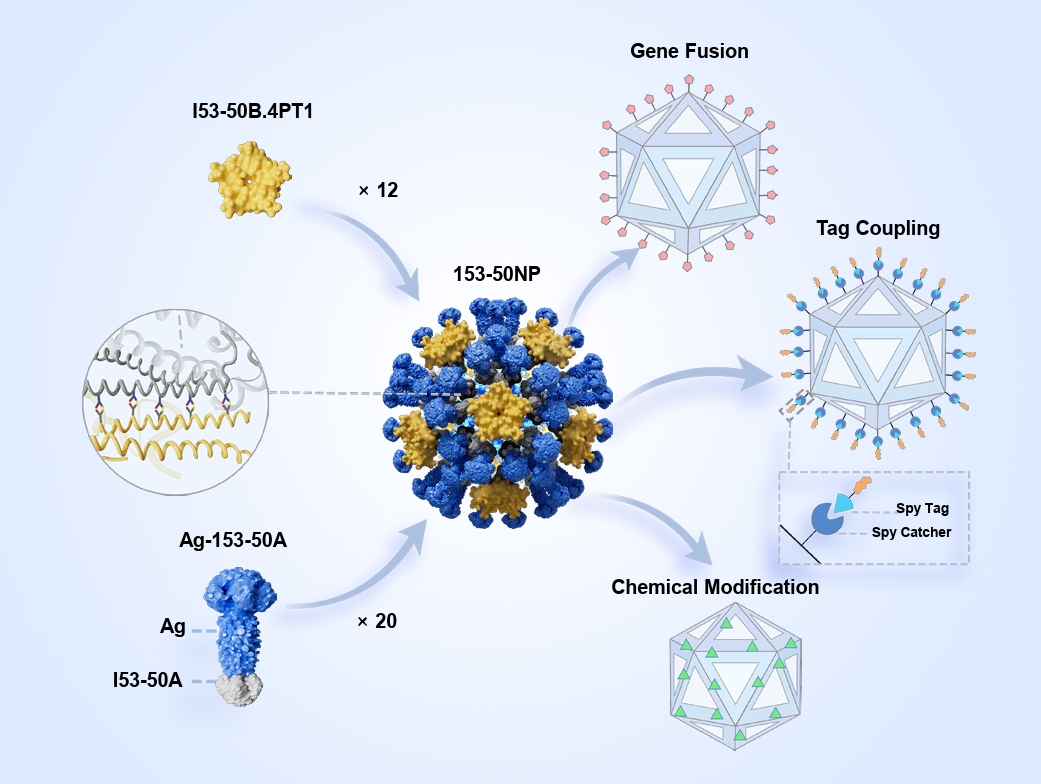
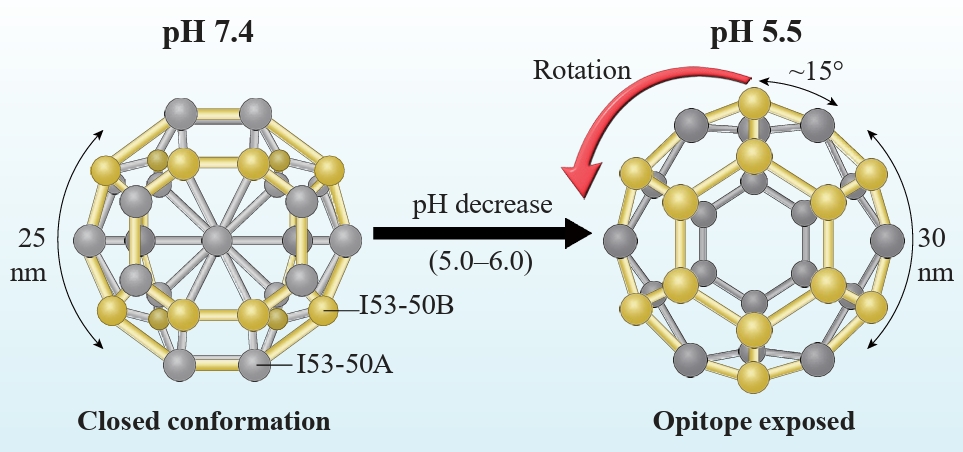
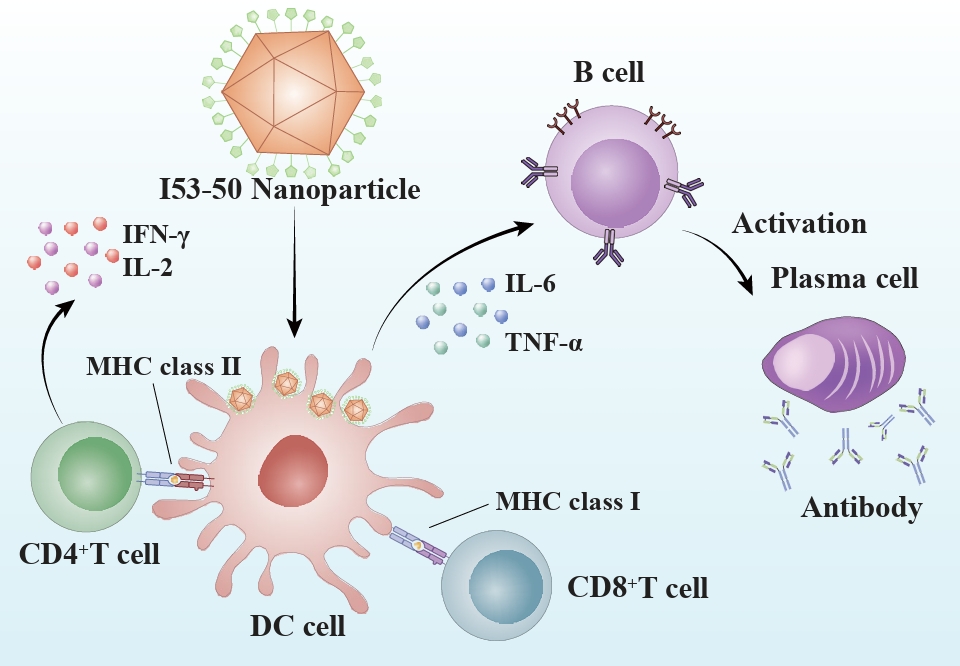
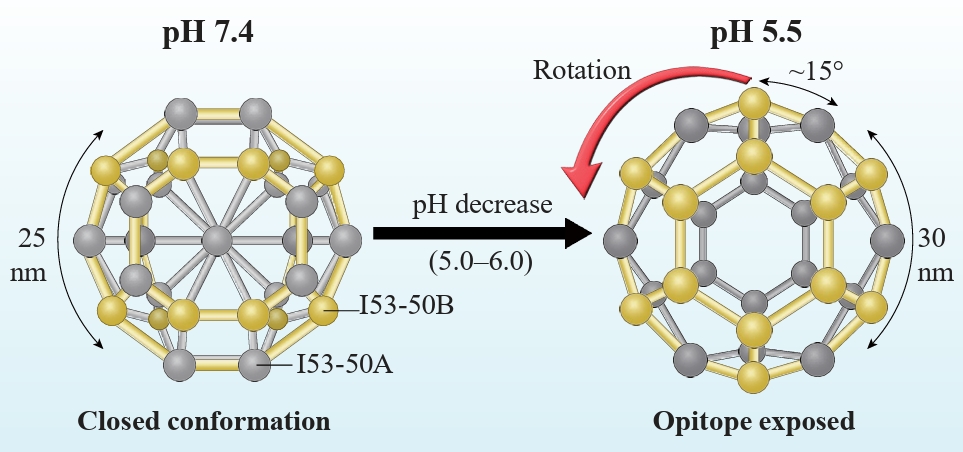
Fig. 1.
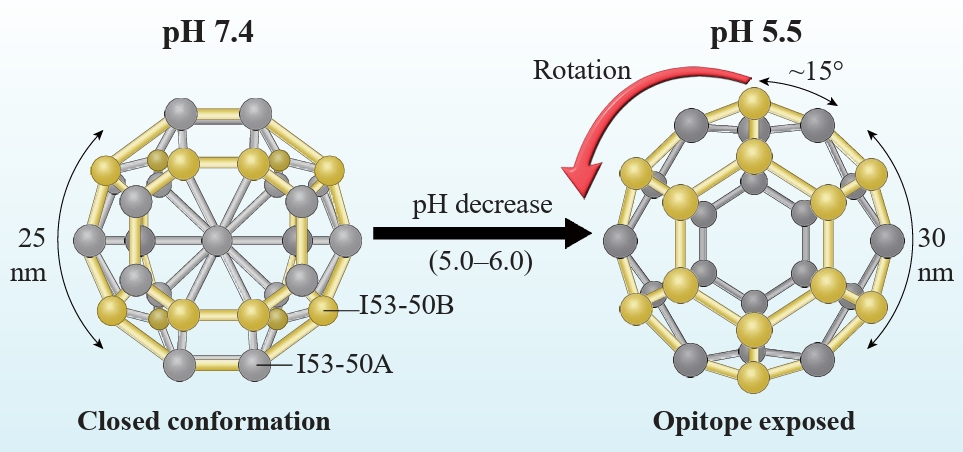
Fig. 2.
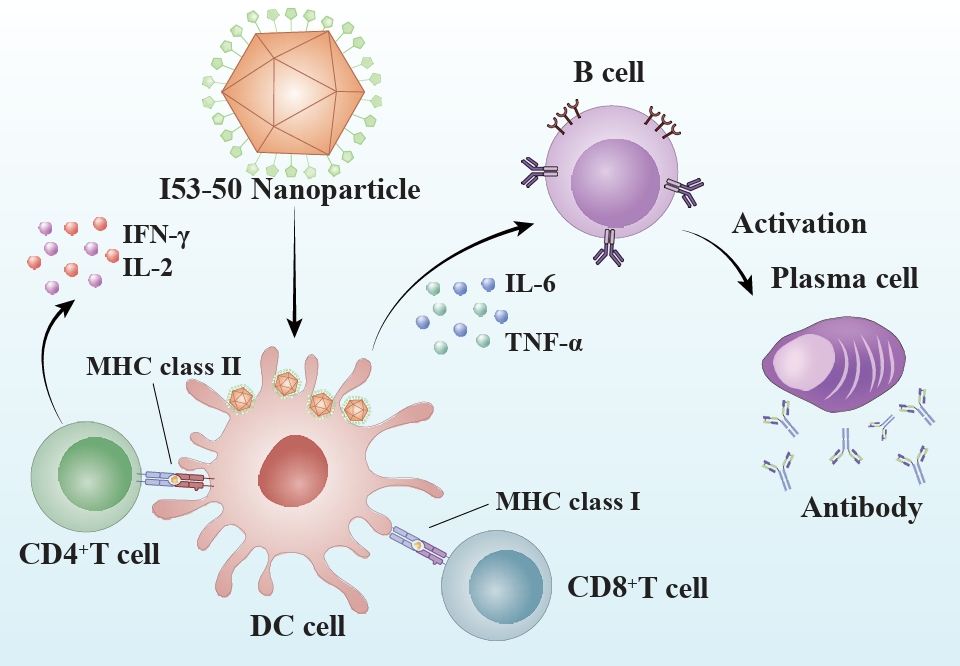
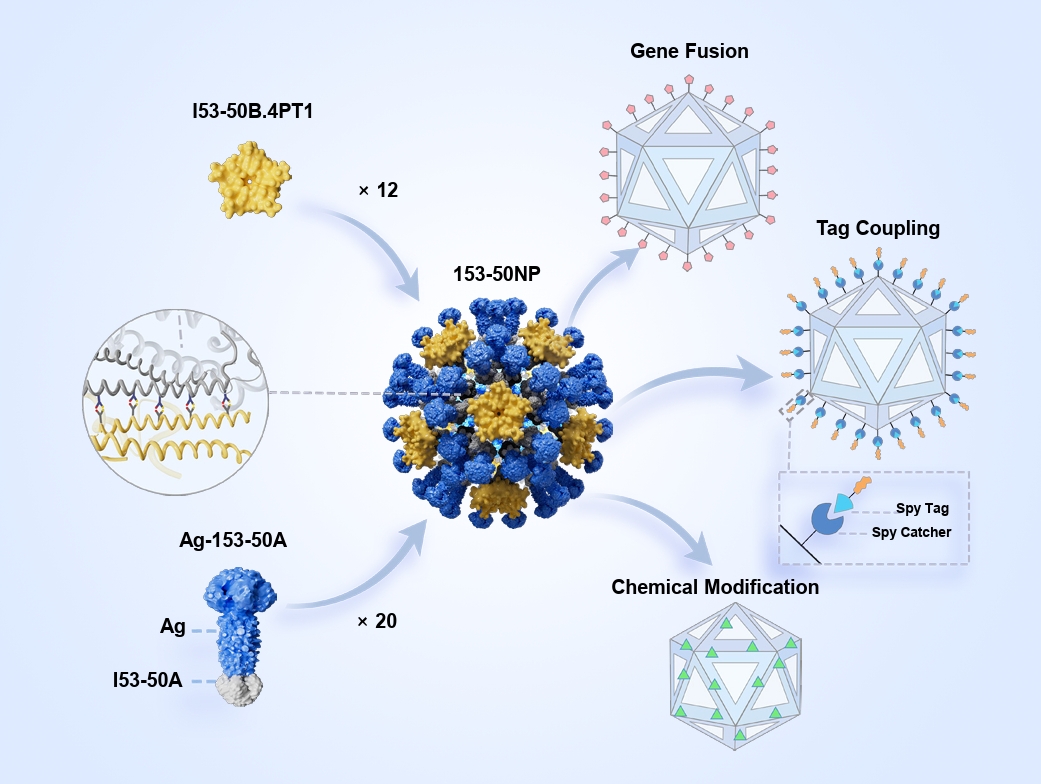
Fig. 3.
| Functionalization strategy | Key characteristics | Advantages | Limitations |
|---|---|---|---|
| Chemical modification | Targeting intrinsic active residues (preferentially N-terminal/C-terminal regions) of I53-50 subunits to conjugate functional groups or molecules via chemical reactions, thereby achieving protein functional modification and enhancement ( |
1. Site-specificity: Enables precise targeting of defined amino acid sites. | 1. Potential structural perturbation: Chemical reactions may affect the native protein conformation and stability. |
| 2. High flexibility: Compatible with a wide range of chemical reagents and reactions. | 2. Risk of heterogeneity: Conjugation reactions can yield heterogeneous products. | ||
| 3. Overcomes spatial constraints: Linker design allows fine-tuning of the distance between the antigen and the nanoparticle surface ( |
3. Dependence on reactive residues: Efficiency is limited by the accessibility of reactive amino acids ( |
||
| Tag coupling | Introducing specific tags into I53-50 subunits via genetic engineering or chemical methods, and achieving site-specific covalent coupling through tag-specific recognition for antigen display, purification, or targeting ( |
1. Stable covalent linkage: Antigen is covalently and stably integrated into the assembly. | 1. Insert size limitation: Large antigen inserts may impair subunit folding and self-assembly. |
| 2. High-density & Uniform display: Enables precise, ordered presentation of 60-240 antigen copies per particle. | 2. Design complexity: Requires optimization of linkers (flexible/rigid) to balance antigen presentation and structural integrity. | ||
| 3. Streamlined production: Single-step expression and self-assembly simplify manufacturing ( |
3. Long redesign cycles: New antigens require de novo gene construction and validation ( |
||
| Gene fusion | Fusing the coding gene of the target antigen with the modified I53-50 subunit gene via recombinant DNA technology, allowing the fusion protein to self-assemble into functional nanoparticles with antigen display ( |
1. Modularity & Versatility: "Plug-and-play" platform for rapid interchange of different SpyTag-fused antigens. | 1. Requires tag engineering: Both carrier and antigen require genetic modification, which may affect their native properties. |
| 2. Site-specific & Efficient: Forms irreversible isopeptide bonds under mild conditions, ensuring high coupling efficiency and homogeneity. | 2. Potential tag immunogenicity: The tags themselves could elicit immune responses (typically low). | ||
| 3. Enhanced stability: Covalent linkage improves complex stability during storage and in vivo ( |
3. Additional conjugation step: Involves an in vitro coupling step compared to genetic fusion ( |
| Parameter / Feature | I53-50NP | VLP | Ferritin | Mi3 |
|---|---|---|---|---|
| Subunit composition | Composed of 24 trimeric (I53-50A) and 12 pentameric (I53-50B) subunits assembling with strict T = 1 icosahedral symmetry ( |
Typically assembles from identical subunits via quasi-equivalent T = 3 or T = 4 icosahedral symmetry, leading to potential structural variability ( |
24 subunits self-assemble into a cage with octahedral (O) symmetry ( |
Trimeric subunits assemble into 2D/3D lattices or finite particles with D3 (dihedral) symmetry ( |
| Thermal stability (Tm) | Exceptionally high. Melting temperature (Tm) > 95°C (for variant I53-50A.1NT1) and remains intact after 1-h incubation at 70°C ( |
Moderate to high. Varies by source; e.g., HBcAg VLP Tm ~70–75°C, but may exhibit kinetic instability ( |
High. Human heavy-chain ferritin (HFH) Tm ~75°C, but stability varies across species and is pH-sensitive ( |
High. Designed Mi3 protein exhibits Tm ~77°C ( |
| Assembly mechanism | Strictly controlled co-assembly. A and B components must mix in a precise stoichiometric ratio to form uniform ~50 nm particles. Enables mosaic display of distinct antigens ( |
Condition-dependent self-assembly. Spontaneous assembly under specific conditions (pH, ionic strength) can lead to heterogeneity in size and morphology ( |
pH-dependent assembly/Disassembly. Assembles at neutral/alkaline pH and disassembles at acidic pH. Useful for cargo loading but may raise concerns about in vivo stability in certain microenvironments ( |
Self-assembly into extended arrays or finite particles. Achieving uniform, finite-sized nanoparticles requires precise design control ( |
| Functionalization | Highly flexible & Precise. Supports genetic fusion (to N/C-termini or specific loops of A/B subunits), chemical conjugation, and tag coupling. Allow precise presentation of 60 or 120 antigen copies with controlled orientation ( |
Flexible but potentially disruptive. Common genetic fusion to N/C-termini or the major immunodominant ( |
Primarily N/C-terminal genetic fusion. Due to octahedral symmetry, maximum valency is limited to 24 copies, resulting in lower antigen density ( |
Primarily genetic fusion to specific sites on the trimeric subunit. Its dihedral symmetry results in antigen geometry distinct from icosahedral presentation ( |
| Name of the vaccine | Construction methods | Immunization effects | Type of connection | Assembly efficiency and conditions |
|---|---|---|---|---|
| RBD-I53-50NP ( |
ΔN1-SpyCatcher first binds to I53-50A1.1PT1 and I53-50B.4PT1, and then fuses with SpyTag attached to the C-terminal end of the SARS-CoV-2 RBD, resulting in ΔN1-SpyCatcher-I53-50NP | RBD-I53-50NP showed stronger affinity for the receptor ACE2 and the neutralizing antibody CB6, indicating good affinity for BCR and triggering higher titers of antibody | tag coupling | I53-50A1.1PT1 and I53-50B.4PT1 were incubated in vitro at 250 mM NaCl, 50 mM Tris-HCl (pH 8.0), containing 5% (v/v) glycerol, and purified by size exclusion chromatography (SEC). No free RBD antigen or residual I53-50 dimer subunits were detected, with high subunit assembly and antigen loading conversion rates. |
| SARS-CoV2 S-I53-50NP ( |
SARS-CoV-2 S-I53-50NP formed by fusing the C-terminus of the pre-fused SARS-CoV-2 S protein to the N-terminus of the I53-50A variant of I53-50A.1NT1, followed by incubation with I53-50B.4PT1 | Enhanced activation of SARS-CoV-2 S protein-specific B cells in vitro induces a strong NAb response | gene fusion | SARS-CoV-2 S-I53-50A.1NT1 and I53-50B.4PT1 were mixed in buffer at 4°C or room temperature for several hours under the following conditions: 50 mM Tris (pH 8), 150 mM NaCl, 100 mM L-arginine, and 5% (w/v) sucrose. Assembly efficiency is typically very high (> 90%), yielding monodisperse nanoparticles with highly uniform morphology. |
| RBD-8G/12G/16G-I53-50NP ( |
SARS-CoV-2 RBD was genetically fused to the N-terminus of I53-50A via a linker of 8, 12, and 16 glycine and serine residues, and later fused to I53-50B.4PT1 to form RBD-I53-50NP | RBD-NP immunization induced a robust and long-lasting neutralizing antibody response, eliciting a strong antigen-specific CD4 T-cell response that skillfully neutralized the mutant strain | gene fusion | I53-50A and I53-50B4.PT1 were incubated at room temperature (25–30°C) for 1–2 h in a buffer containing 250 mM NaCl, 50 mM Tris-HCl pH 7.4–8.0, and 5% glycerol. The subunit assembly conversion rate approached 100% (size exclusion chromatography (SEC) showed no free subunit peaks), with a polydispersity index (PDI) < 0.1 (determined by dynamic light scattering (DLS)). |
| E2E1-I53-50NP ( |
The N-terminus of I53-50A1.NT1 is fused to the C-terminus of E2E1, which later fuses with I53-50B.4PT1 to form E2E1-I53-50NP | The number of neutralizing viruses in animals immunized with E2E1-NPs was significantly higher than that in animals receiving monoimmunization, and both the width and potency of Nab were significantly improved, with an increase in potency of up to 80-fold | gene fusion | E2E1-I53-50A and I53-50B.4PT1 were incubated overnight at 4°C in Tris-buffered saline (TBS, pH ~7.5) supplemented with 5% glycerol or L-arginine, achieving assembly efficiencies typically > 90%. Both SEC monomodal profiles and NS-EM images indicate the product consists of highly uniform protein nanoparticles, with no detectable free protein or aggregates. |
| p67C-I53-50NP ( |
ECF p67C is linked to the I53-50A N-terminus and the p67C-I53-50A fusion gene formed is fused to I53-50B.4PT1 to form p67c-I53-50NP | p67C-I53-50 immunized animals produced high p67C-specific IgG1 and IgG2 antibody titers as well as strong CD4+ T-cell responses, with the highest amount of IFN-γ secretion | gene fusion | p67C‑I53‑50A and I53-50B.4PT1 were incubated at room temperature (RT) for 30 min to 2 h in 50 mM Tris‑HCl, pH 8.0, 250–500 mM NaCl, supplemented with 50 mM glycine or 0.75% CHAPS, at room temperature (RT) for 30 min to 2 h. Size exclusion chromatography (SEC) revealed a single symmetric peak for the assembled nanoparticles, indicating a homogeneous product. Dynamic light scattering (DLS) showed a uniform particle size distribution consistent with the expected dimensions. |
| gB-I53-50-NP ( |
HCV viral gB gene fused to the N-terminus of I53-50A1 via C-terminal fusion, forming gB-I53-50A1 fused to I53-50B.4PT1 to form gB-I53-50NP | Induction of serum polyclonal antibodies by gB-I53-50NP vaccine in monkeys significantly protects humanized mice against EBV infection and lymphomagenesis, and the protective effect persists for a long period of time up to 10 weeks after immunization of monkeys | gene fusion | I53-50A1 and I53-50B4.PT1 can be efficiently assembled by incubating them for 1 h in assembly buffer (250 mM NaCl, 50 mM Tris pH 8.0, 5% glycerol). |
| SOSIP-I53-50NP ( |
HIV viral SOSIP Env trimer fused to I53-50A to form the SOSIP-I53-50A construct, which later fused to I53-50B.4PT1 to form SOSIP-I53-50NP | SOSIP-I53-50 NP in immunized animals, binds antibodies with reactive IgG titers up to 50–100 μg/ml, neutralizes viruses with titers ID50 > 100, Abs targets a wide range of neutralizing epitopes. | gene fusion | SOSIP-I53-50A and I53-50B.4PT1 were incubated overnight at 4°C in 25 mM Tris-HCl (pH 8.0), 500 mM NaCl, 5% glycerol at 4°C overnight. Dynamic light scattering analysis revealed the hydrodynamic radius of the assembled nanoparticles ranged from 246 to 292 Å with a low polydispersity index, indicating uniform and monodisperse particle size distribution. |
| DS-Cav1-I53-50 and Sc9-10-I53-50 ( |
The extracellular domain sequences of DS-Cav1 or Sc9-10 were fused with the I53-50A subunit gene to construct the DS-Cav1-I53-50A and Sc9-10-I53-50A fusion protein. The fusion protein was secreted and expressed in a stable Chinese hamster ovary (CHO) cell line, while the I53-50B subunit was expressed in Escherichia coli BL21 (DE3). The two subunits self-assembled in a 3:1 mass ratio to form DS-Cav1-I53-50 and Sc9-10-I53-50 nanoparticles. | In mouse models, total IgG titers, D25 competitive neutralization antibody titers, and live virus neutralization antibody titers against RSV A2 strain were elevated. Due to the antigenic properties of Sc9-10, Sc9-10-I53-50 exhibited slightly higher immunogenicity than DS-Cav1-I53-50. | gene fusion | DS-Cav1-I53-50A and Sc9-10-I53-50A were mixed with I53-50B in a buffer system of 50 mM Tris-HCl, 300 mM NaCl, 0.75% CHAPS, pH 7.4, and incubated at room temperature (RT) for 2 h. Purified by size exclusion chromatography (SEC), the particles exhibited uniform size and high assembly efficiency. |
| DS2-I53-50 ( |
The DS2 antigen gene was directly fused with the I53-50A gene to construct the DS2-I53-50A fusion protein. The DS2-I53-50A fusion protein was transiently expressed in Expi293F mammalian cells, while the I53-50B subunit was induced in Escherichia coli BL21(DE3). Following separate purifications, the two subunits self-assembled in a 1:1 molar ratio to form stable DS2-I53-50 nanoparticles. | Following immunization of BALB/c mice, the induced DS2-specific IgG titer was 2.9 times that of free DS2, with the IgG1/IgG2a ratio decreasing to 2.6, indicating a Th1-biased response. This significantly promoted the proliferation of germinal center B cells (CD19⁺B220⁺CD95⁺GL-7⁺) and follicular helper T cells (CD4⁺CXCR5⁺PD-1⁺), activated dendritic cells and macrophages, and expanded the CD4⁺ central/ effector memory T cell populations. | gene fusion | Mix DS2-I53-50A with I53-50B in PBS (containing 5% glycerol) and incubate at 25°C for 3 h. Size exclusion chromatography shows that all particles assembled successfully, with the most uniform chromatographic peak observed for I53-50. |
Table 1.
Table 2.
Table 3.
TOP
 MSK
MSK
 ePub Link
ePub Link Cite this Article
Cite this Article