- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- For Contributors
- Policies
- E-Submission

- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- Policies
- For Contributors
Articles
- Page Path
- HOME > J. Microbiol > Volume 64(4); 2026 > Article
-
Full article
Inhibitory effects of acetyl-11-keto-β-boswellic acid (AKBA) on human cytomegalovirus (HCMV) in vitro - Bingquan Chu1, Zhiwei Ding1, Xinna Wu2, Yunchuang Chang2, Chunxia Wu2, Yicheng Fu2, Genxiang Mao2,*, Sanying Wang2,*
-
Journal of Microbiology 2026;64(4):e2601007.
DOI: https://doi.org/10.71150/jm.2601007
Published online: March 25, 2026
1School of Biological and Chemical Engineering, Zhejiang University of Science and Technology, Hangzhou 310023, P. R. China
2Zhejiang Key Laboratory of Geriatrics and Geriatrics Institute of Zhejiang Province, Zhejiang Hospital, Hangzhou 310030, P. R. China
- *Correspondence Genxiang Mao maogenxiang@zju.edu.cn Sanying Wang sanyingwang309@126.com
© The Microbiological Society of Korea
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
- 967 Views
- 20 Download
ABSTRACT
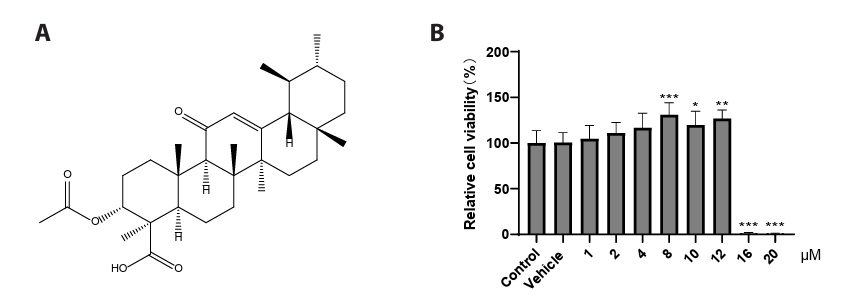
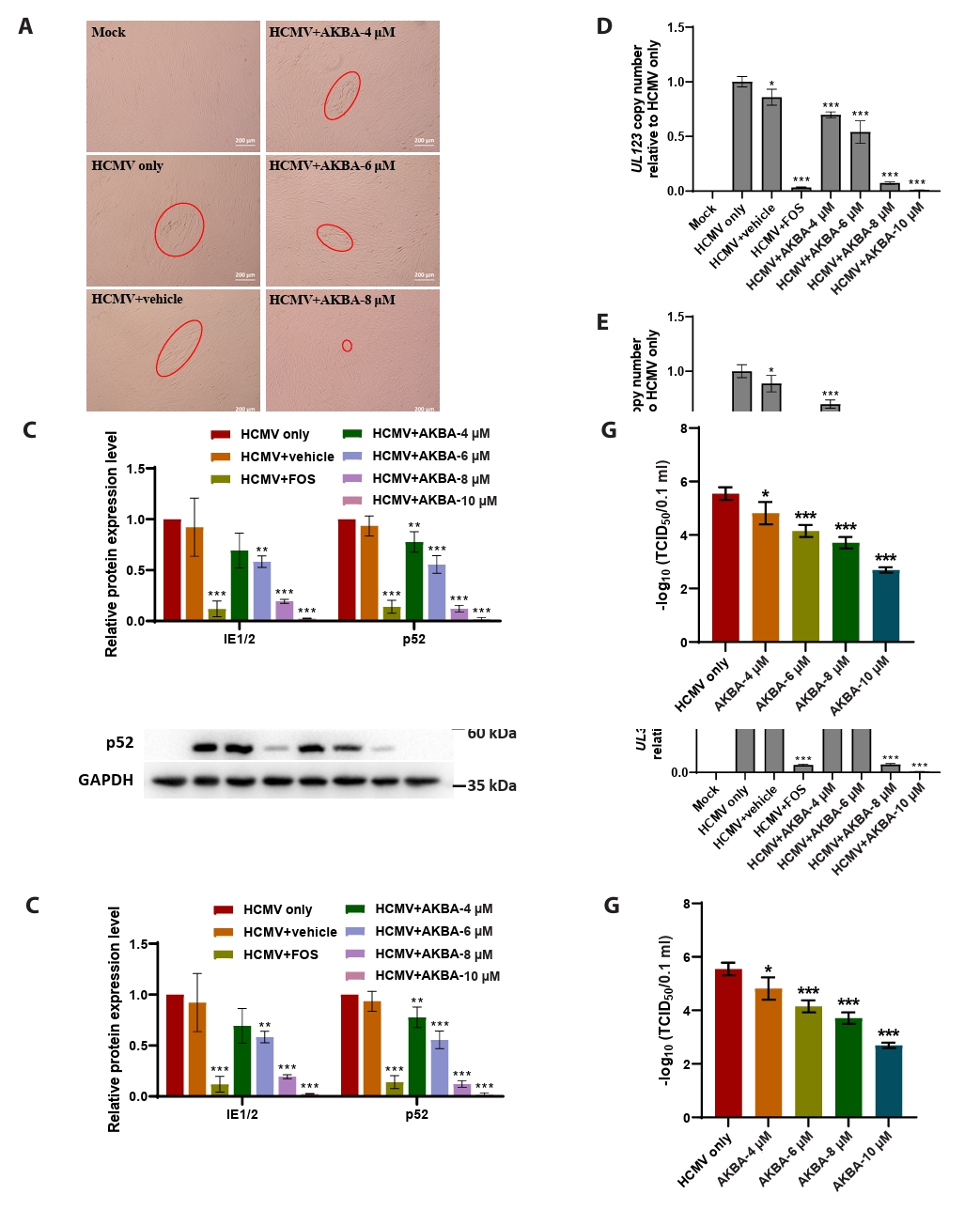
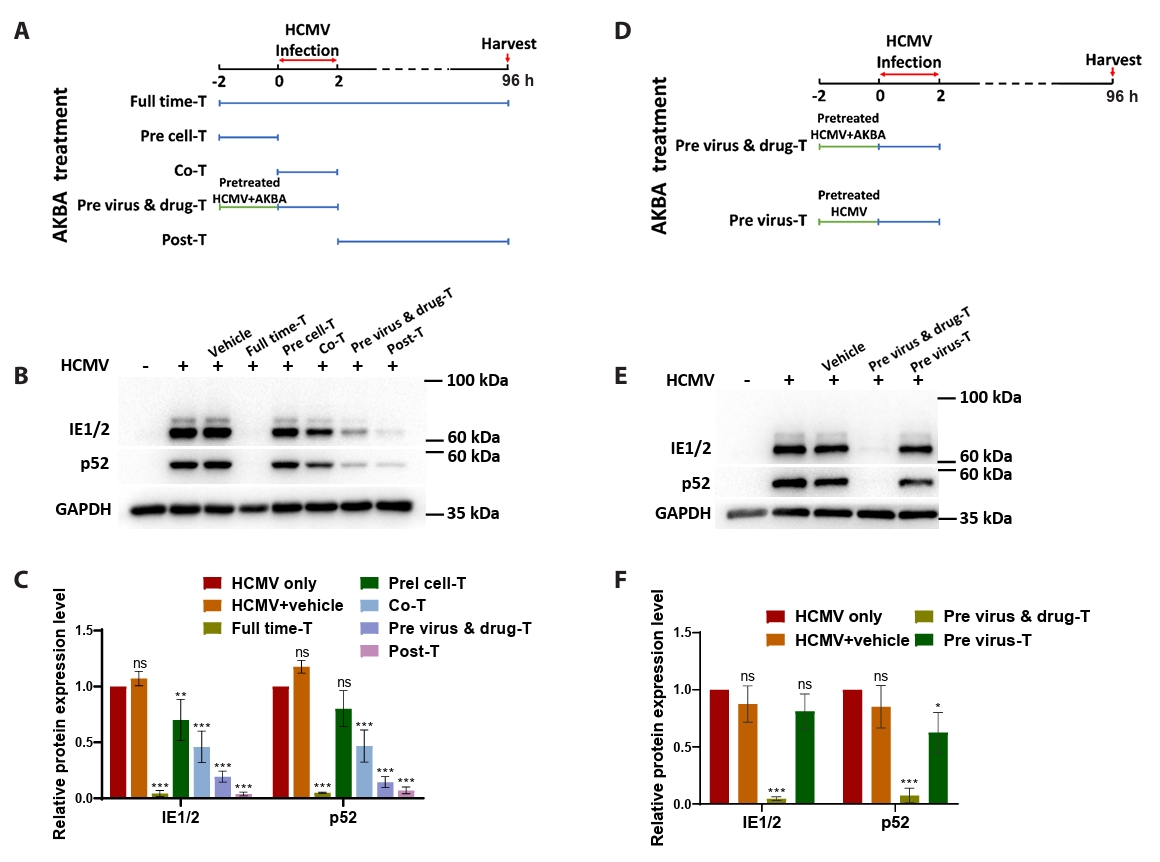
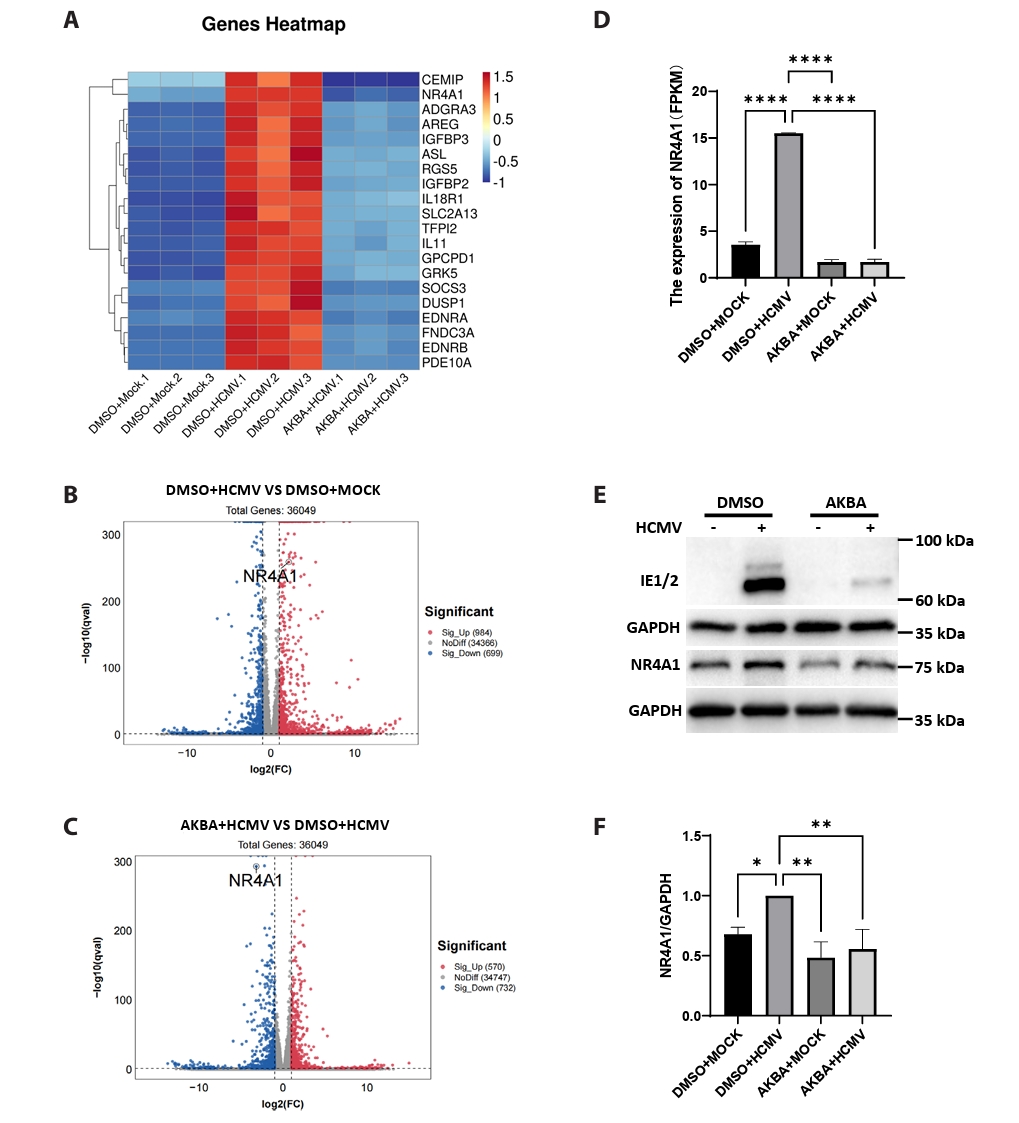
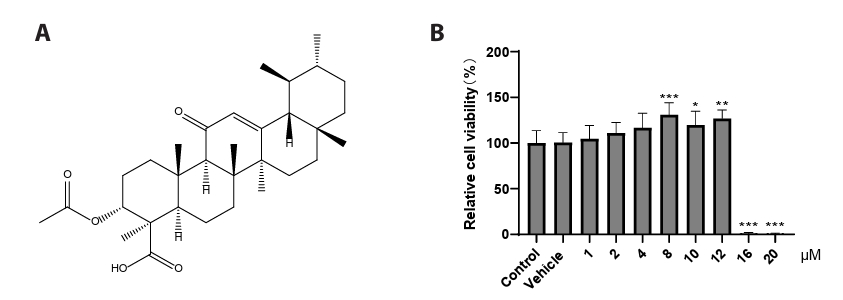
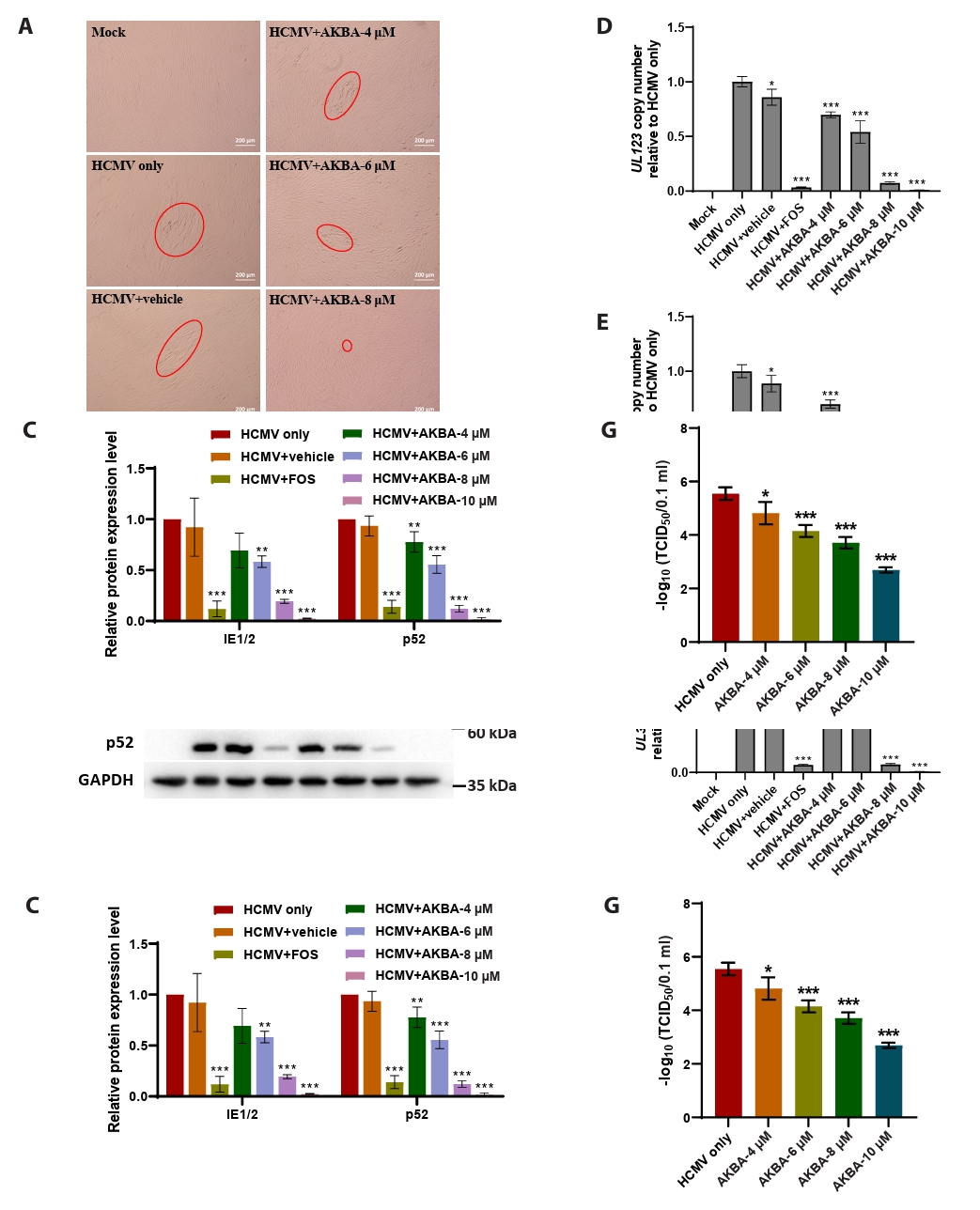
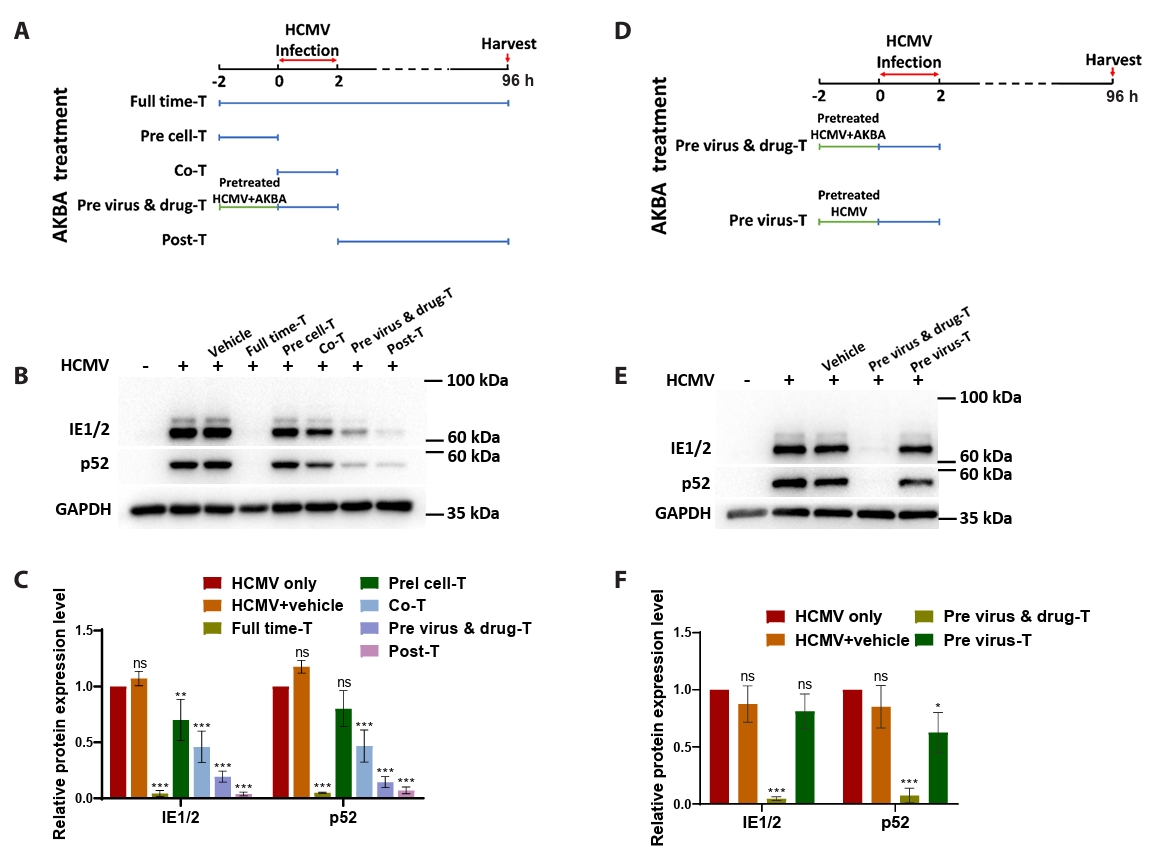
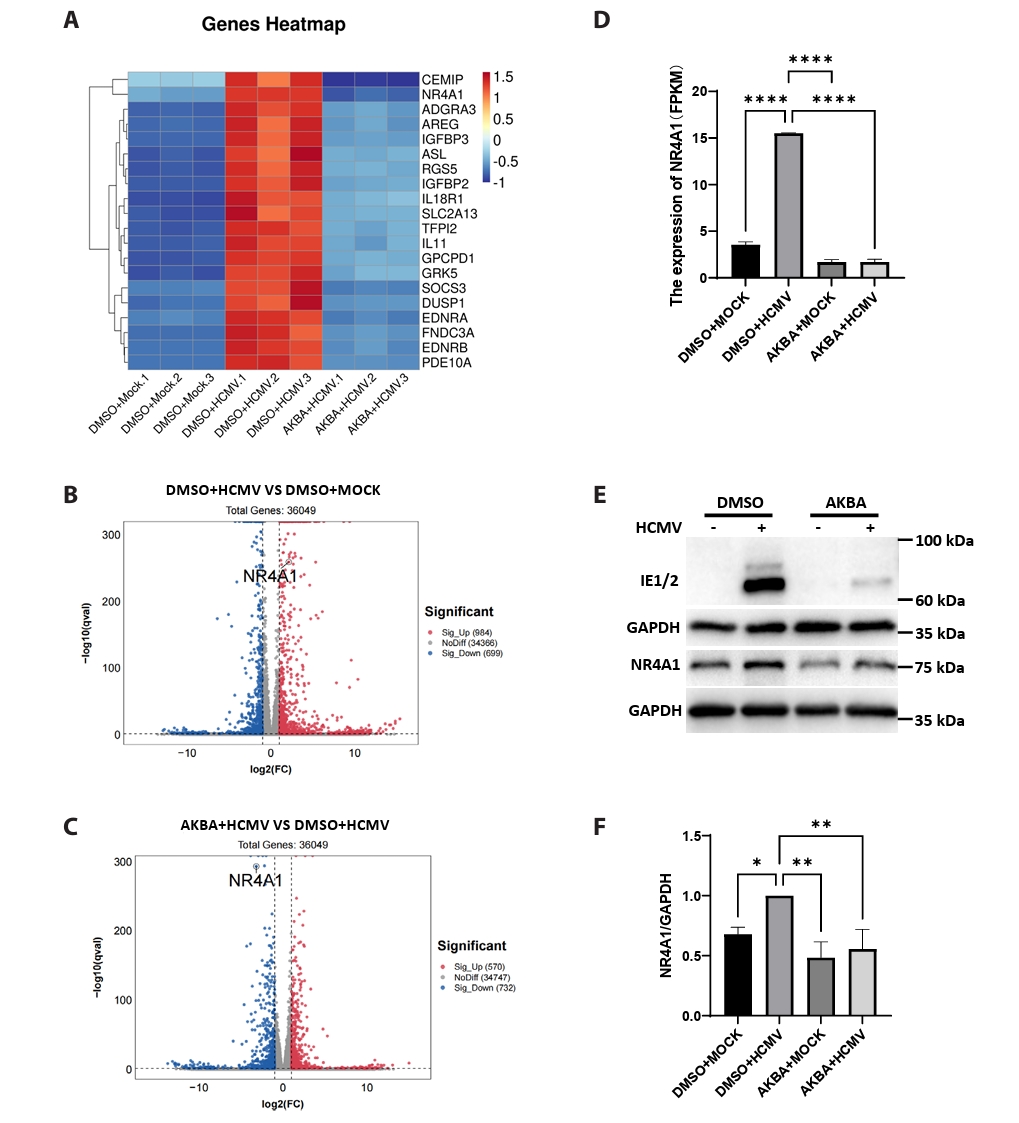
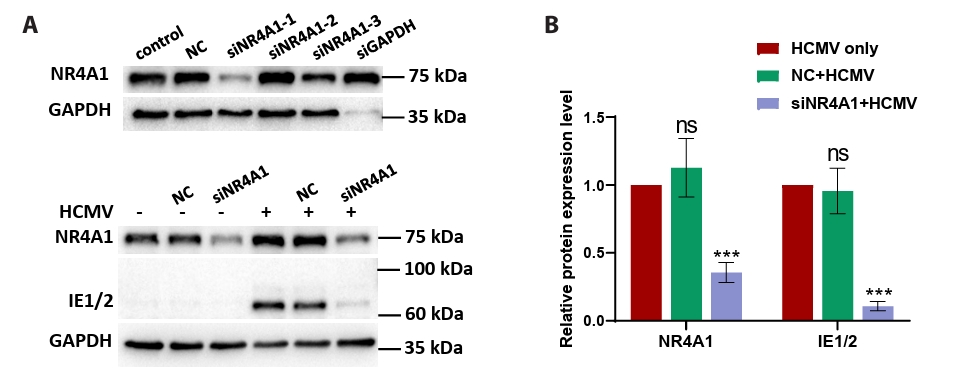
- This study presents the first investigation of acetyl-11-keto-β-boswellic acid (AKBA)’s anti-human cytomegalovirus (HCMV) activity in vitro and elucidates its underlying mechanisms. In HCMV Towne strain-infected WI-38 cells, AKBA (1-12 μM) exhibited negligible cytotoxicity while significantly suppressing virus-induced cytopathic effects (CPE) at 6–10 μM, with dose-dependent reduction of viral proteins (IE1/2 and p52) expression, viral DNA copy number (UL123, UL44, and UL32), and infectious viral progeny titer (TCID50). Time-of-addition experiments demonstrated the primary antiviral activity of AKBA during post-entry phase, along with direct virion inactivation. Transcriptome analysis revealed that AKBA significantly downregulated the expression of the host factor NR4A1 induced by HCMV, a finding further validated by Western blotting. Further gene knockdown experiments confirmed that silencing NR4A1 significantly reduced the expression of viral proteins IE1/2, thereby validating NR4A1 as a key host factor for HCMV infection. These findings indicate that AKBA has a potent and dose-dependent inhibitory effect on HCMV replication in WI-38 cells, and proves that this effect is mediated through two different mechanisms: one is the downregulation of the expression of the key host factor NR4A1, and the other is the direct inactivation of HCMV viral particles.
Introduction
Materials and Methods
Results
Discussion
Acknowledgments
This work was supported by Joint TCM Science & Technology Projects of National Demonstration Zones for Comprehensive TCM Reform (GZY-KJS-ZJ-2025-074 and GZY-ZJ-KJ-24055), and TCM Key Laboratory Cultivation Base of Zhejiang Province for the Development and Clinical Transformation of Immunomodulatory Drugs (TCM-202402).
Conflict of Interest
The authors have no conflict of interest to report.
Ethical Statements
None.
Supplementary Information
Fig. S1.
- Atanasoff KE, Ophir SI, Parsons AJ, Paredes Casado J, Lurain NS, et al. 2025. N-arylpyrimidinamine (NAPA) compounds are broadly acting inhibitors of human cytomegalovirus infection and spread. Antivir Res. 233: 106044.ArticlePubMed
- Bradley E, Poole E, Reeves MB. 2024. The triterpenoid MOMORDIN-Ic inhibits HCMV by preventing the initiation of gene expression in eukaryotic cells. Pathogens. 13: 546.ArticlePubMedPMC
- Britt WJ. 2018. Maternal immunity and the natural history of congenital human cytomegalovirus infection. Viruses. 10: 405.ArticlePubMedPMC
- Chen M, Hu J, Zhou X, Gao M, Li N, et al. 2025. Long non-coding RNA THRIL promotes influenza virus replication by inhibiting the antiviral innate immune response. Viruses. 17: 153.ArticlePubMedPMC
- Chen SJ, Wang SC, Chen YC. 2022. Challenges, recent advances and perspectives in the treatment of human cytomegalovirus infections. Trop Med Infect Dis. 7: 439.ArticlePubMedPMC
- Crough T, Khanna R. 2009. Immunobiology of human cytomegalovirus: from bench to bedside. Clin Microbiol Rev. 22: 76–98. ArticlePubMedPMCLink
- García-Ramírez JJ, Ruchti F, Huang H, Simmen K, Angulo A, et al. 2001. Dominance of virus over host factors in cross-species activation of human cytomegalovirus early gene expression. J Virol. 75: 26–35. ArticlePubMedPMCLink
- Gong Y, Jiang X, Yang S, Huang Y, Hong J, et al. 2022. The biological activity of 3-O-acetyl-11-keto-β-boswellic acid in nervous system diseases. Neuromolecular Med. 24: 374–384. ArticlePubMedPMCPDF
- Gourin C, Alain S, Hantz S. 2023. Anti-CMV therapy, what next? A systematic review. Front Microbiol. 14: 1321116.ArticlePubMedPMC
- Griffiths P, Reeves M. 2021. Pathogenesis of human cytomegalovirus in the immunocompromised host. Nat Rev Microbiol. 19: 759–773. ArticlePubMedPMCPDF
- Han Q, Qian Y, Bai L, Zhou J, Hao Y, et al. 2025. Injectable nano-micron AKBA delivery platform for treatment of tendinopathy in a rat model. J Biomed Mater Res A. 113: e37844. ArticlePubMed
- He P, Sun J, Tan J, Shi YN, Zhang CJ, et al. 2025. From physicochemical constraints to clinical prospects of celastrol: challenges and nano delivery strategies. Int J Nanomedicine. 20: 10907–10931. ArticlePubMedPMCLink
- Isomura H, Stinski MF. 2013. Coordination of late gene transcription of human cytomegalovirus with viral DNA synthesis: recombinant viruses as potential therapeutic vaccine candidates. Expert Opin Ther Targets. 17: 157–166. ArticlePubMed
- Kalejta RF. 2008. Tegument proteins of human cytomegalovirus. Microbiol Mol Biol Rev. 72: 249–265. ArticlePubMedPMCLink
- Kosolapov D, Jac P, Riasova P, Pouskova J, Polasek M, et al. 2025. Advances and challenges in the analysis of boswellic acids by separation methods. Crit Rev Anal Chem. 55: 748–774. ArticlePubMed
- Lauß J, Kappacher C, Isser O, Huck CW, Rainer M. 2024. Species-specific quantification of bioactive boswellic acids in Boswellia resin using NIR spectroscopy, HPLC and Multivariate data analysis. Spectrochim Acta A Mol Biomol Spectrosc. 316: 124384.ArticlePubMed
- Levrier A, Soudier P, Garenne D, Izri Z, Bowden S, et al. 2025. A synthetic cell phage cycle. Nat Commun. 17: 557.ArticlePubMedPMCPDF
- Li Z, Pang J, Dong L, Yu X. 2021. Structural basis for genome packaging, retention, and ejection in human cytomegalovirus. Nat Commun. 12: 4538.ArticlePubMedPMCPDF
- Liu TW, Zhao YM, Jin KY, Wang JX, Zhao XF. 2024. KAT8 is upregulated and recruited to the promoter of Atg8 by FOXO to induce H4 acetylation for autophagy under 20-hydroxyecdysone regulation. J Biol Chem. 300: 105704.ArticlePubMedPMC
- Mao G, Li H, Ding X, Meng X, Wang G, et al. 2016. Suppressive effects of sirtinol on human cytomegalovirus (hCMV) infection and hCMV-induced activation of molecular mechanisms of senescence and production of reactive oxygen species. Mech Ageing Dev. 158: 62–69. ArticlePubMedPMC
- Messinger CJ, Lipsitch M, Bateman BT, He M, Huybrechts KF, et al. 2020. Association between congenital cytomegalovirus and the prevalence at birth of microcephaly in the United States. JAMA Pediatr. 174: 1159–1167. ArticlePubMedPMC
- Nischang V, Witt FM, Borner F, Gomez M, Jordan PM, et al. 2023. Frankincense preparation promotes formation of inflammation-resolving lipid mediators by manipulating lipoxygenases in human innate immune cells. Front Pharmacol. 14: 1332628.ArticlePubMed
- Rand U, Kubsch T, Kasmapour B, Cicin-Sain L. 2020. A novel triple-fluorescent HCMV strain reveals gene expression dynamics and anti-herpesviral drug mechanisms. Front Cell Infect Microbiol. 10: 536150.ArticlePubMed
- Rozman B, Nachshon A, Levi Samia R, Lavi M, Schwartz M, et al. 2022. Temporal dynamics of HCMV gene expression in lytic and latent infections. Cell Rep. 39: 110653.ArticlePubMedPMC
- Safe S, Shrestha R, Mohankumar K. 2021. Orphan nuclear receptor 4A1 (NR4A1) and novel ligands. Essays Biochem. 65: 877–886. ArticlePubMedPMCPDF
- Senaweera S, Edwards TC, Kankanala J, Wang Y, Sahani RL, et al. 2022. Discovery of N-benzyl hydroxypyridone carboxamides as a novel and potent antiviral chemotype against human cytomegalovirus (HCMV). Acta Pharm Sin B. 12: 1671–1684. ArticlePubMed
- Shirvanimoghaddam K, Akbari MK, Yadav R, Al-Tamimi AK, Naebe M. 2021. Fight against COVID-19: The case of antiviral surfaces. APL Mater. 9: 031112.ArticlePubMedPMCPDF
- Tommasi C, Rogerson C, Depledge DP, Jones M, Naeem AS, et al. 2020. Kallikrein-mediated cytokeratin 10 degradation is required for varicella zoster virus propagation in skin. J Invest Dermatol. 140: 774–784. ArticlePubMed
- Van Damme E, Van Loock M. 2014. Functional annotation of human cytomegalovirus gene products: an update. Front Microbiol. 5: 218.ArticlePubMedPMC
- Wan MSM, Muhammad R, Koliopoulos MG, Roumeliotis TI, Choudhary JS, et al. 2023. Mechanism of assembly, activation and lysine selection by the SIN3B histone deacetylase complex. Nat Commun. 14: 2556.ArticlePubMedPMCPDF
- Wang SY, Zhang J, Xu XG, Su HL, Xing WM, et al. 2020. Inhibitory effects of piceatannol on human cytomegalovirus (hCMV) in vitro. J Microbiol. 58: 716–723. ArticlePubMedPDF
- Wild M, Karner D, Eickhoff J, Wagner S, Kicuntod J, et al. 2023. Combined treatment with host-directed and anticytomegaloviral kinase inhibitors: mechanisms, synergisms and drug resistance barriers. Pharmaceutics. 15: 2680.ArticlePubMedPMC
- Yao X, Jing T, Geng Q, Pang M, Zhao X, et al. 2023. Dual analysis of wild-type and attenuated Orf virus and host cell transcriptomes revealed novel virus-host cell interactions. mSphere. 8: e00398-23. ArticlePubMedPMCLink
- Zhang T, Potgieter TI, Kosche E, Ruckert J, Ostermann E, et al. 2023. Thioxothiazolo[3,4-a]quinazoline derivatives inhibit the human cytomegalovirus alkaline nuclease. Antivir Res. 217: 105696.ArticlePubMed
- Zhang SM, Rehling D, Jemth AS, Throup A, Landazuri N, et al. 2021. NUDT15-mediated hydrolysis limits the efficacy of anti-HCMV drug ganciclovir. Cell Chem Biol. 28: 1693–1702. ArticlePubMed
- Zuhair M, Smit GSA, Wallis G, Jabbar F, Smith C, et al. 2019. Estimation of the worldwide seroprevalence of cytomegalovirus: A systematic review and meta-analysis. Rev Med Virol. 29: e2034. ArticlePubMedLink
References
Supplementary Information
References
Citations

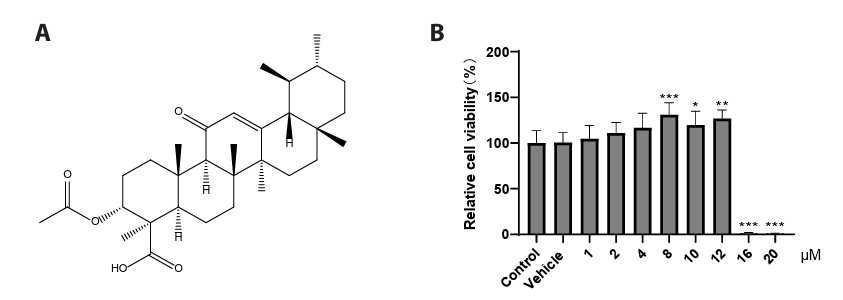
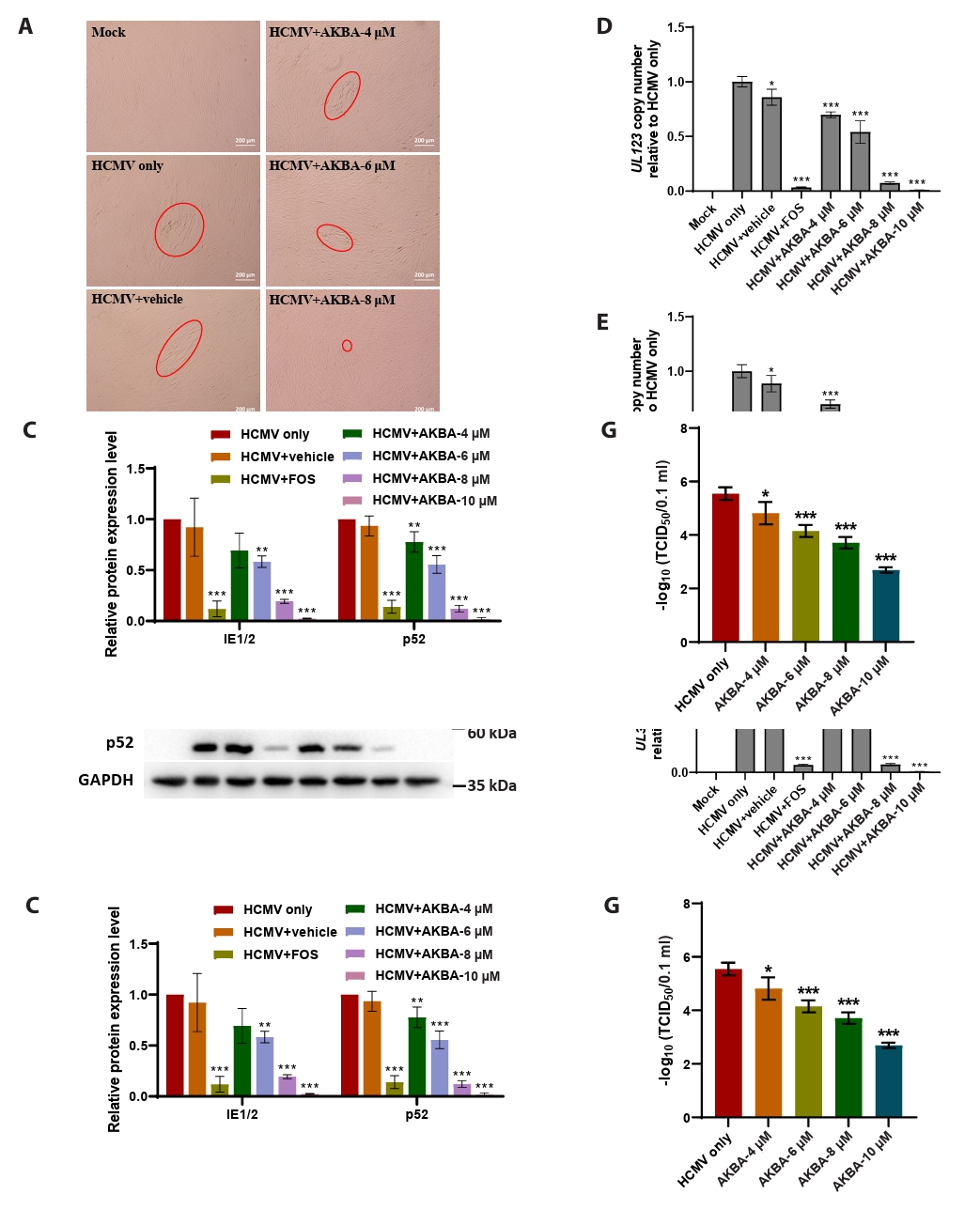
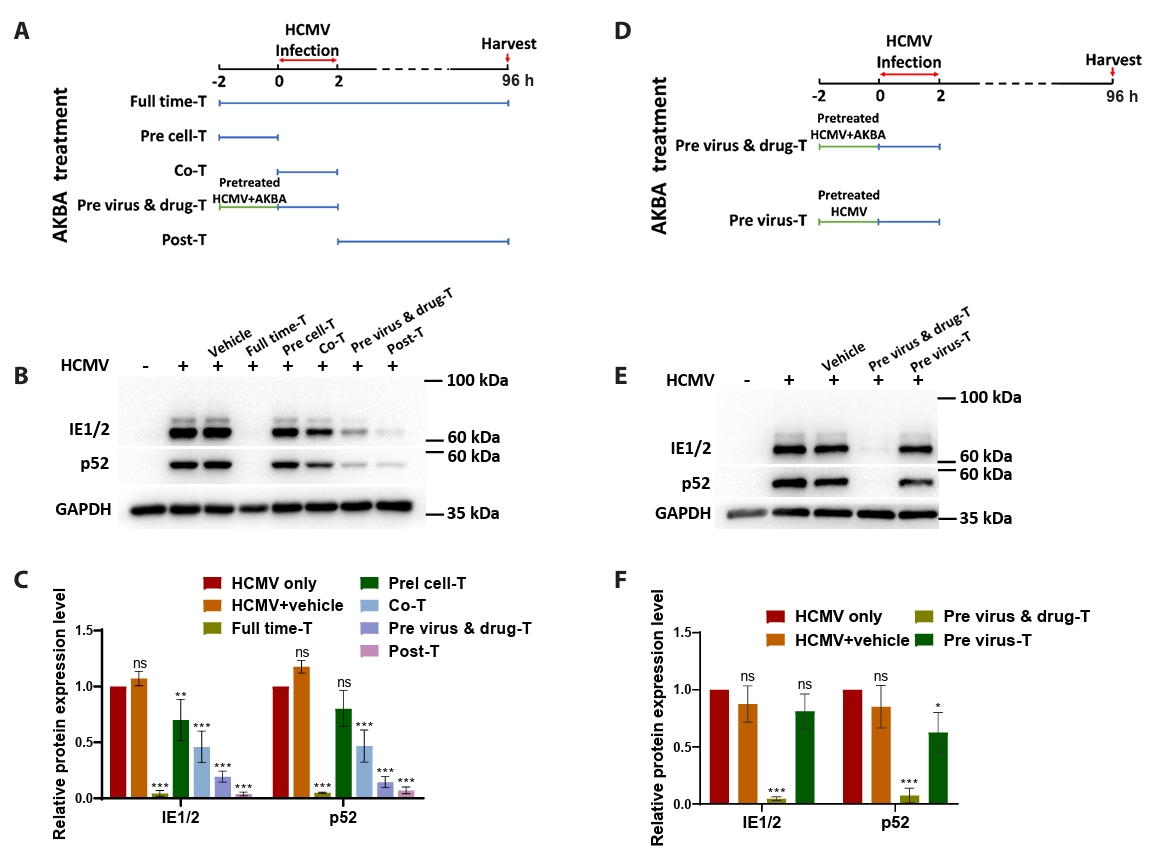
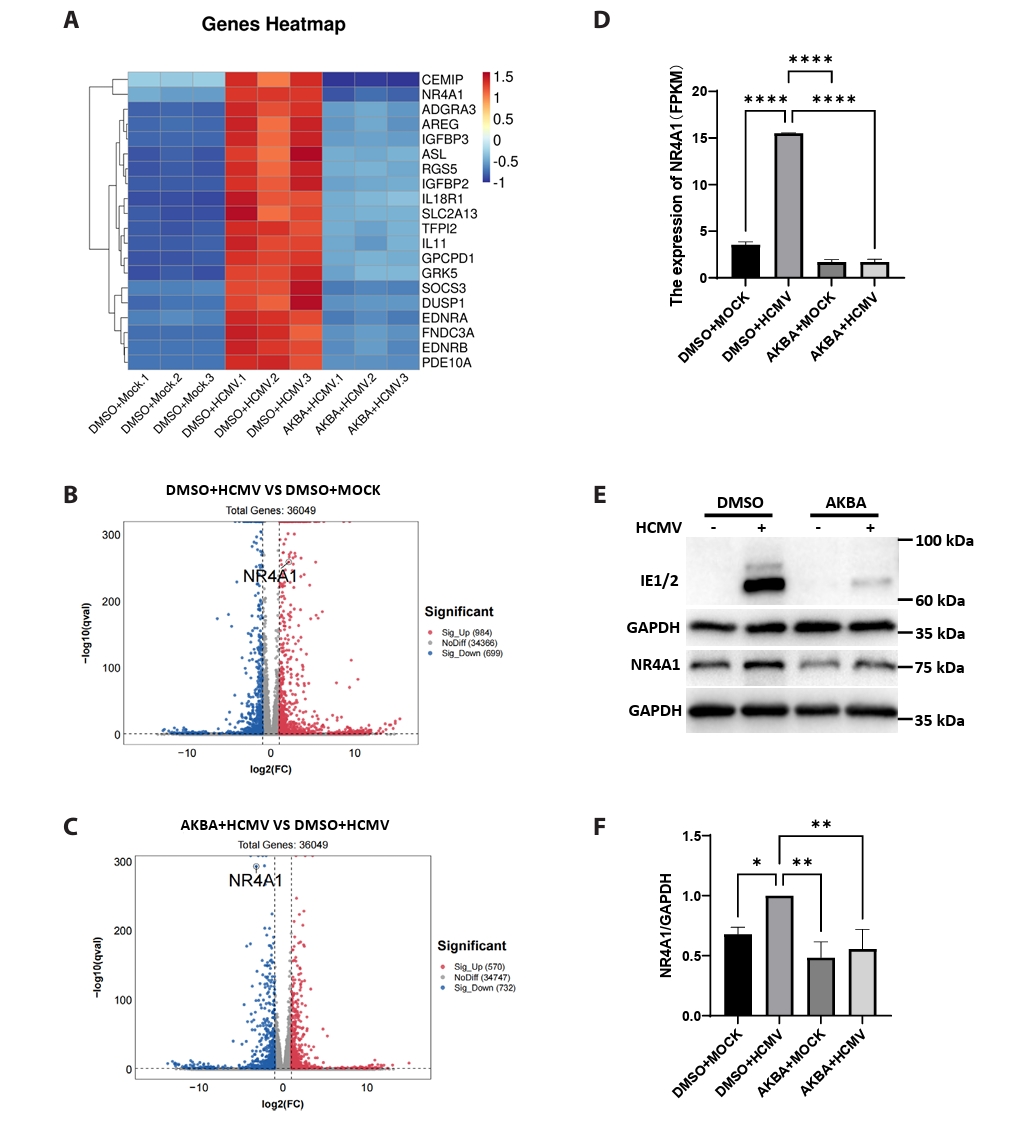
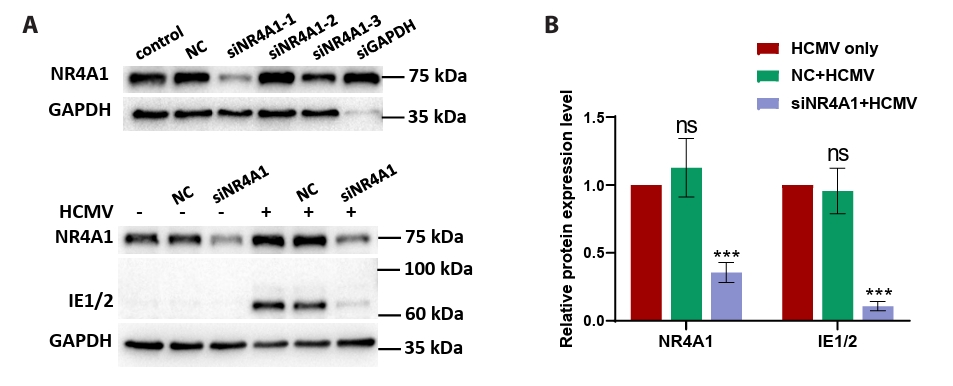
Fig. 1.
Fig. 2.
Fig. 3.
Fig. 4.
Fig. 5.
| Primer name | Primer sequence (5'-3') |
|---|---|
| UL123-F | TCTGCCAGGACATCTTTCTC |
| UL123-R | GTGACCAAGGCCACGACGTT |
| UL44-F | ACTGCCGTGCACGTTGCGTA |
| UL44-R | ACTTGCCGCTGTTCCCGACG |
| UL32-F | GGTTTCTGGCTCGTGGATGTCG |
| UL32-R | CACACAACACCGTCGTCCGATTAC |
| GAPDH-F | CTGTTGCTGTAGCCAAATTCGT |
| GAPDH-R | ACCCACTCCTCCACCTTTGAC |
| siRNA | siRNA sequence (5'-3') |
|---|---|
| siNR4A1-1-Sense | GCACCUUCAUGGACGGCUA dTdT |
| siNR4A1-1-Antisense | UAGCCGUCCAUGAAGGUGC dTdT |
| siNR4A1-2-Sense | CCUUCAAGUUCGAGGACUU dTdT |
| siNR4A1-2-Antisense | AAGUCCUCGAACUUGAAGG dTdT |
| siNR4A1-3-Sense | UGGUGAAGGAAGUUGUCCGAA dTdT |
| siNR4A1-3-Antisense | UUCGGACAACUUCCUUCACCA dTdT |
Table 1.
Table 2.
TOP
 MSK
MSK
 ePub Link
ePub Link Cite this Article
Cite this Article